diff --git a/.github/workflows/tests.yml b/.github/workflows/tests.yml
index cac3d2d..ad7aae3 100644
--- a/.github/workflows/tests.yml
+++ b/.github/workflows/tests.yml
@@ -1,59 +1,59 @@
name: tests
on:
- push:
- branches: [main]
- pull_request:
+ push:
+ branches: [main]
+ pull_request:
concurrency:
- group: tests-${{ github.workflow }}-${{ github.ref }}
- cancel-in-progress: true
+ group: tests-${{ github.workflow }}-${{ github.ref }}
+ cancel-in-progress: true
jobs:
- pre-commit:
- runs-on: ubuntu-latest
- steps:
- - uses: actions/checkout@v4
- - uses: actions/setup-python@v5
- with:
- python-version: '3.10'
- - uses: pre-commit/action@v3.0.1
-
- build:
- runs-on: ubuntu-latest
- steps:
- - name: Check out the repository
- uses: actions/checkout@v4
- with:
- fetch-depth: 2
-
- - name: Install uv
- uses: astral-sh/setup-uv@v4
-
- - name: Set up Python
- run: uv python install
-
- - name: Install dependencies
- run: uv sync --all-extras --dev
-
- - name: Install Quarto
- uses: quarto-dev/quarto-actions/setup@v2
- with:
- version: "1.5.57"
-
- - name: set timezone
- run: |
- TZ="Europe/London" &&
- sudo ln -snf /usr/share/zoneinfo/$TZ /etc/localtime
-
- - name: install linux deps
- run: |
- sudo apt-get -y install openssl graphviz nano texlive graphviz-dev unzip build-essential
-
- - name: build the book
- run: |
- uv run quarto render --execute
-
- - name: success
- run: |
- echo "Success in building book without errors!"
+ pre-commit:
+ runs-on: ubuntu-latest
+ steps:
+ - uses: actions/checkout@v4
+ - uses: actions/setup-python@v5
+ with:
+ python-version: "3.10"
+ - uses: pre-commit/action@v3.0.1
+
+ build:
+ runs-on: ubuntu-latest
+ steps:
+ - name: Check out the repository
+ uses: actions/checkout@v4
+ with:
+ fetch-depth: 2
+
+ - name: Install uv
+ uses: astral-sh/setup-uv@v4
+
+ - name: Set up Python
+ run: uv python install
+
+ - name: Install dependencies
+ run: uv sync --all-extras --dev
+
+ - name: Install Quarto
+ uses: quarto-dev/quarto-actions/setup@v2
+ with:
+ version: "1.5.57"
+
+ - name: set timezone
+ run: |
+ TZ="Europe/London" &&
+ sudo ln -snf /usr/share/zoneinfo/$TZ /etc/localtime
+
+ - name: install linux deps
+ run: |
+ sudo apt-get -y install openssl graphviz nano texlive graphviz-dev unzip build-essential
+
+ - name: build the book
+ run: |
+ uv run quarto render --execute
+
+ - name: success
+ run: |
+ echo "Success in building book without errors!"
diff --git a/data-transform.ipynb b/data-transform.ipynb
index 55f2df7..c20ac1c 100644
--- a/data-transform.ipynb
+++ b/data-transform.ipynb
@@ -11,7 +11,7 @@
"\n",
"It's very rare that data arrive in exactly the right form you need. Often, you'll need to create some new variables or summaries, or maybe you just want to rename the variables or reorder the observations to make the data a little easier to work with.\n",
"\n",
- "You'll learn how to do all that (and more!) in this chapter, which will introduce you to data transformation using the **pandas** package and a new dataset on flights that departed New York City in 2013.\n",
+ "You'll learn how to do all that (and more!) in this chapter, which will introduce you to data transformation using the **polars** package and a new dataset on flights that departed New York City in 2013.\n",
"\n",
"The goal of this chapter is to give you an overview of all the key tools for transforming a data frame, a special kind of object that holds tabular data.\n",
"\n",
@@ -19,7 +19,7 @@
"\n",
"### Prerequisites\n",
"\n",
- "In this chapter we'll focus on the **pandas** package, one of the most widely used tools for data science. You'll need to ensure you have **pandas** installed. To do this, you can run"
+ "In this chapter we'll focus on the **polars** package, one of the most widely used tools for data science. You'll need to ensure you have **polars** installed. To do this, you can run\n"
]
},
{
@@ -29,7 +29,7 @@
"metadata": {},
"outputs": [],
"source": [
- "import pandas as pd"
+ "import polars as pl"
]
},
{
@@ -37,9 +37,9 @@
"id": "438cc0a4",
"metadata": {},
"source": [
- "If this command fails, you don't have **pandas** installed. Open up the terminal in Visual Studio Code (Terminal -> New Terminal), `cd` to the folder you are working in, and type in `uv add pandas`.\n",
+ "If this command fails, you don't have **polars** installed. Open up the terminal in Visual Studio Code (Terminal -> New Terminal), `cd` to the folder you are working in, and type in `uv add polars`.\n",
"\n",
- "Furthermore, if you wish to check which version of **pandas** you're using, it's"
+ "Furthermore, if you wish to check which version of **polars** you're using, it's\n"
]
},
{
@@ -49,7 +49,7 @@
"metadata": {},
"outputs": [],
"source": [
- "pd.__version__"
+ "pl.__version__"
]
},
{
@@ -57,11 +57,11 @@
"id": "0c5e5b82",
"metadata": {},
"source": [
- "You'll also need the data. Most of the time, data will need to be loaded from a file or the internet. These data are no different, but one of the amazing things about **pandas** is how many different types of data it can load, including from files on the internet.\n",
+ "You'll also need the data. Most of the time, data will need to be loaded from a file or the internet. These data are no different, but one of the amazing things about **polars** is how many different types of data it can load, including from files on the internet.\n",
"\n",
"The data is around 50MB in size so you will need a good internet connection or a little patience for it to download.\n",
"\n",
- "Let's download the data:"
+ "Let's download the data:\n"
]
},
{
@@ -71,8 +71,20 @@
"metadata": {},
"outputs": [],
"source": [
+ "import io\n",
+ "\n",
+ "import requests\n",
+ "\n",
"url = \"https://raw.githubusercontent.com/byuidatascience/data4python4ds/master/data-raw/flights/flights.csv\"\n",
- "flights = pd.read_csv(url)"
+ "resp = requests.get(url, timeout=60)\n",
+ "resp.raise_for_status()\n",
+ "\n",
+ "flights = pl.read_csv(\n",
+ " io.BytesIO(resp.content),\n",
+ " null_values=[\"NA\"],\n",
+ " truncate_ragged_lines=True,\n",
+ " ignore_errors=True,\n",
+ ")"
]
},
{
@@ -80,7 +92,7 @@
"id": "2907635c",
"metadata": {},
"source": [
- "If the above code worked, then you've downloaded the data in CSV format and put it in a data frame. Let's look at the first few rows using the `.head()` function that works on all **pandas** data frames."
+ "If the above code worked, then you've downloaded the data in CSV format and put it in a data frame. Let's look at the first few rows using the `.head()` function that works on all **polars** data frames.\n"
]
},
{
@@ -98,7 +110,7 @@
"id": "68aada55",
"metadata": {},
"source": [
- "To get more general information on the columns, the data types (`dtypes`) of the columns, and the size of the dataset, use `.info()`."
+ "To get more general information on the columns, the data types (`dtypes`) of the columns, and the size of the dataset, use `.glimpse()`.\n"
]
},
{
@@ -108,7 +120,7 @@
"metadata": {},
"outputs": [],
"source": [
- "flights.info()"
+ "flights.glimpse(max_items_per_column=5)"
]
},
{
@@ -116,22 +128,23 @@
"id": "100189b8",
"metadata": {},
"source": [
- "You might have noticed the short abbreviations that appear in the `Dtypes` column. These tell you the type of the values in their respective columns: `int64` is short for integer (eg whole numbers) and `float64` is short for double-precision floating point number (these are real numbers). `object` is a bit of a catch all category for any data type that **pandas** is not really confident about inferring. Although not found here, other data types include `string` for text and `datetime` for combinations of a date and time.\n",
+ "You might have noticed the short abbreviations that appear in the `dtype` column. These tell you the type of the values in their respective columns: `i64` is short for integer (eg whole numbers) and `f64` is short for double-precision floating point number (these are real numbers). **polars** has an `object` data type which allows storing arbitrary Python objects, but this makes you lose performance benefits, as **polars** is **strictly typed**. Although not found here, other data types include `str` for text and `datetime` for combinations of a date and time.\n",
"\n",
"The table below gives some of the most common data types you are likely to encounter.\n",
"\n",
- "| **Name of data type** | **Type of data** |\n",
- "|:----------:|:-------------:|\n",
- "| float64 | real numbers |\n",
- "| category | categories |\n",
- "| datetime64 | date times |\n",
- "| int64 | integers |\n",
- "| bool | True or False |\n",
- "| string | text |\n",
+ "| **Name of data type** | **Type of data** |\n",
+ "| :-------------------: | :--------------: |\n",
+ "| Float64 | real numbers |\n",
+ "| Categorical | categories |\n",
+ "| Datetime | date and time |\n",
+ "| Date | date |\n",
+ "| Int64 | integers |\n",
+ "| Boolean | True or False |\n",
+ "| String | text |\n",
"\n",
"The different column data types are important because the operations you can perform on a column depend so much on its \"type\"; for example, you can remove all punctuation from strings while you can multiply ints and floats.\n",
"\n",
- "We would like to work with the `\"time_hour\"` variable in the form of a datetime; fortunately, **pandas** makes it easy to perform that conversion on that specific column"
+ "We would like to work with the `\"time_hour\"` variable in the form of a datetime; fortunately, **polars** makes it easy to perform that conversion on that specific column\n"
]
},
{
@@ -141,7 +154,7 @@
"metadata": {},
"outputs": [],
"source": [
- "flights[\"time_hour\"]"
+ "flights.get_column(\"time_hour\")"
]
},
{
@@ -151,7 +164,7 @@
"metadata": {},
"outputs": [],
"source": [
- "flights[\"time_hour\"] = pd.to_datetime(flights[\"time_hour\"], format=\"%Y-%m-%dT%H:%M:%SZ\")"
+ "flights.with_columns(pl.col(\"time_hour\").str.to_datetime())"
]
},
{
@@ -159,17 +172,17 @@
"id": "6dc43cee",
"metadata": {},
"source": [
- "## **pandas** basics\n",
+ "## **polars** basics\n",
"\n",
- "**pandas** is a really comprehensive package, and this book will barely scratch the surface of what it can do. But it's built around a few simple ideas that, once they've clicked, make life a lot easier.\n",
+ "**polars** is a really comprehensive package, and this book will barely scratch the surface of what it can do. But it's built around a few simple ideas that, once they've clicked, make life a lot easier.\n",
"\n",
- "Let’s start with the absolute basics. The most basic pandas object is DataFrame. A DataFrame is a 2-dimensional data structure that can store data of different types (including characters, integers, floating point values, categorical data, even lists) in columns. It is made up of rows and columns (with each row-column cell containing a value), plus two bits of contextual information: the index (which carries information about each row) and the column names (which carry information about each column).\n",
+ "Let’s start with the absolute basics. The most basic polars object is DataFrame. A DataFrame is a 2-dimensional data structure that can store data of different types (including characters, integers, floating point values, categorical data, even lists) in columns. It is made up of rows and columns (with each row-column cell containing a value), plus contextual information: column names (which carry information about each column).\n",
"\n",
- "\n",
- "\n",
- "Perhaps the most important notion to have about **pandas** data frames is that they are built around an index that sits on the left-hand side of the data frame. Every time you perform an operation on a data frame, you need to think about how it might or might not affect the index; or, put another way, whether you want to modify the index.\n",
+ "::: {.callout-note}\n",
+ "Note: If you're coming from **pandas**, be aware that **polars** does not use an index column and each row is indexed by its integer position in the table.\n",
+ ":::\n",
"\n",
- "Let's see a simple example of this with a made-up data frame:"
+ "\n"
]
},
{
@@ -179,7 +192,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df = pd.DataFrame(\n",
+ "df = pl.DataFrame(\n",
" data={\n",
" \"col0\": [0, 0, 0, 0],\n",
" \"col1\": [0, 0, 0, 0],\n",
@@ -187,7 +200,6 @@
" \"col3\": [\"a\", \"b\", \"b\", \"a\"],\n",
" \"col4\": [\"alpha\", \"gamma\", \"gamma\", \"gamma\"],\n",
" },\n",
- " index=[\"row\" + str(i) for i in range(4)],\n",
")\n",
"df.head()"
]
@@ -197,7 +209,7 @@
"id": "185ba56e",
"metadata": {},
"source": [
- "You can see there are 5 columns (named `\"col0\"` to `\"col4\"`) and that the index consists of four entries named `\"row0\"` to `\"row3\"`."
+ "You can see there are 5 columns (named `\"col0\"` to `\"col4\"`).\n"
]
},
{
@@ -205,14 +217,14 @@
"id": "3f325661",
"metadata": {},
"source": [
- "A second key point you should know is that the operations on a **pandas** data frame can be chained together. We need not perform one assignment per line of code; we can actually do multiple assignments in a single command.\n",
+ "A second key point you should know is that the operations on a **polars** data frame can be chained together. We need not perform one assignment per line of code; we can actually do multiple assignments in a single command.\n",
"\n",
"Let's see an example of this. We're going to string together four operations:\n",
"\n",
- "1. we will use `query()` to find only the rows where the destination `\"dest\"` column has the value `\"IAH\"`. This doesn't change the index, it only removes irrelevant rows. In effect, this step removes rows we're not interested in.\n",
- "2. we will use `groupby()` to group rows by the year, month, and day (we pass a list of columns to the `groupby()` function). This step changes the index; the new index will have three columns in that track the year, month, and day. In effect, this step changes the index.\n",
- "3. we will choose which columns we wish to keep after the `groupby()` operation by passing a list of them to a set of square brackets (the double brackets are because it's a list within a data frame). Here we just want one column, `\"arr_delay\"`. This doesn't affect the index. In effect, this step removes columns we're not interested in.\n",
- "4. finally, we must specify what `groupby()` operation we wish to apply; when aggregating the information in multiple rows down to one row, we need to say how that information should be aggregated. In this case, we'll use the `mean()`. In effect, this step applies a statistic to the variable(s) we selected earlier, across the groups we created earlier."
+ "1. we will use `filter()` to find only the rows where the destination `\"dest\"` column has the value `\"IAH\"`. We use the `pl.col()` expression to select the column for filtering condition. In effect, this step removes rows we're not interested in.\n",
+ "2. we will use `group_by()` to group rows by the year, month, and day (we pass a list of columns to the `group_by()` function).\n",
+ "3. we will choose which columns to perform aggregation on after the `group_by()` operation by using the `pl.col()` expression inside `agg()`, to select the column. Here we just want one column, `\"arr_delay\"`. In effect, this step removes columns we're not interested in.\n",
+ "4. finally, we must specify what `agg()` operation we wish to apply; when aggregating the information in multiple rows down to one row, we need to say how that information should be aggregated. In this case, we'll use the `mean()`. In effect, this step applies a statistic to the variable(s) we selected earlier, across the groups we created earlier.\n"
]
},
{
@@ -222,7 +234,9 @@
"metadata": {},
"outputs": [],
"source": [
- "(flights.query(\"dest == 'IAH'\").groupby([\"year\", \"month\", \"day\"])[[\"arr_delay\"]].mean())"
+ "flights.filter(pl.col(\"dest\") == \"IAH\").group_by([\"year\", \"month\", \"day\"]).agg(\n",
+ " pl.col(\"arr_delay\").mean()\n",
+ ")"
]
},
{
@@ -230,16 +244,15 @@
"id": "b8b85551",
"metadata": {},
"source": [
- "You can see here that we've created a new data frame with a new index. To do it, we used four key operations:\n",
+ "You can see here that we've created a new data frame. To do it, we used three key operations:\n",
"\n",
"1. manipulating rows\n",
- "2. manipulating the index\n",
- "3. manipulating columns\n",
- "4. applying statistics\n",
+ "2. manipulating columns\n",
+ "3. applying statistics\n",
"\n",
"Most operations you could want to do to a single data frame are covered by these, but there are different options for each of them depending on what you need.\n",
"\n",
- "Let's now dig a bit more into these operations."
+ "Let's now dig a bit more into these operations.\n"
]
},
{
@@ -249,7 +262,7 @@
"source": [
"## Manipulating Rows in Data Frames\n",
"\n",
- "Let's create some fake data to show how this works."
+ "Let's create some fake data to show how this works.\n"
]
},
{
@@ -261,13 +274,13 @@
"source": [
"import numpy as np\n",
"\n",
- "df = pd.DataFrame(\n",
+ "df = pl.DataFrame(\n",
" data=np.reshape(range(36), (6, 6)),\n",
- " index=[\"a\", \"b\", \"c\", \"d\", \"e\", \"f\"],\n",
- " columns=[\"col\" + str(i) for i in range(6)],\n",
- " dtype=float,\n",
+ " schema=[\"col\" + str(i) for i in range(6)],\n",
+ ")\n",
+ "df.insert_column(\n",
+ " 6, pl.Series(\"col6\", [\"apple\", \"orange\", \"pineapple\", \"mango\", \"kiwi\", \"lemon\"])\n",
")\n",
- "df[\"col6\"] = [\"apple\", \"orange\", \"pineapple\", \"mango\", \"kiwi\", \"lemon\"]\n",
"df"
]
},
@@ -278,9 +291,10 @@
"source": [
"### Accessing Rows\n",
"\n",
- "To access a particular row directly, you can use `df.loc['rowname']` or `df.loc[['rowname1', 'rowname2']]` for two different rows.\n",
+ "To access a particular row directly, you can get that by index (location in the data) or predicate/expression using `.row()`, which returns as a tuple.\n",
+ "Remember that Python indices begin from zero, so to retrieve the first row by index you would use `.row(0)`:\n",
"\n",
- "For example,"
+ "For example,\n"
]
},
{
@@ -290,7 +304,11 @@
"metadata": {},
"outputs": [],
"source": [
- "df.loc[[\"a\", \"b\"]]"
+ "# Gets the first row of the DataFrame\n",
+ "df.row(0)\n",
+ "\n",
+ "# Gets the fifth row of the DataFrame\n",
+ "df.row(4)"
]
},
{
@@ -298,7 +316,7 @@
"id": "18124edd",
"metadata": {},
"source": [
- "But you can also access particular rows based on their location in the data frame using `.iloc`. Remember that Python indices begin from zero, so to retrieve the first row you would use `.iloc[0]`:\n"
+ "We can also access particular rows based on a predicate using `.row()` with the `by_predicate` parameter.\n"
]
},
{
@@ -308,7 +326,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df.iloc[0]"
+ "df.row(by_predicate=pl.col(\"col6\") == \"mango\")"
]
},
{
@@ -316,7 +334,7 @@
"id": "ca822472",
"metadata": {},
"source": [
- "This works for multiple rows too. Let's grab the first and third rows (in positions 0 and 2) by passing a list of positions:"
+ "To get the row as a dictionary instead of a tuple with a mapping of column names to row values, specify `named=True`\n"
]
},
{
@@ -326,15 +344,29 @@
"metadata": {},
"outputs": [],
"source": [
- "df.iloc[[0, 2]]"
+ "# Get the first row of the DataFrame as a dictionary\n",
+ "df.row(0, named=True)\n",
+ "\n",
+ "# Get the row where col6 is \"mango\" as a dictionary\n",
+ "df.row(by_predicate=pl.col(\"col6\") == \"mango\", named=True)"
]
},
{
"cell_type": "markdown",
- "id": "381eb34d",
+ "id": "980b7be6",
+ "metadata": {},
+ "source": [
+ "We can also access rows using the `.slice()` function. As the function name implies, we get a slice of the DataFrame. we can use this to get a single row or a number of rows. To use this, we give it an offset - a start index, negative indexing is supported to index from the bottom of the DataFrame - and a length of the slice. This returns a DataFrame\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "f7e8e892",
"metadata": {},
+ "outputs": [],
"source": [
- "There are other ways to access multiple rows that make use of *slicing* but we'll leave that topic for another time."
+ "df.slice(-2, 2)"
]
},
{
@@ -342,9 +374,9 @@
"id": "77f67ac2",
"metadata": {},
"source": [
- "### Filtering rows with query\n",
+ "### Filtering rows\n",
"\n",
- "As with the flights example, we can also filter rows based on a condition using `query()`:"
+ "As with the flights example, we can also filter rows based on a condition using `filter()`:\n"
]
},
{
@@ -354,7 +386,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df.query(\"col6 == 'kiwi' or col6 == 'pineapple'\")"
+ "df.filter((pl.col(\"col6\") == \"kiwi\") | (pl.col(\"col6\") == \"pineapple\"))"
]
},
{
@@ -362,7 +394,7 @@
"id": "000eb292",
"metadata": {},
"source": [
- "For numbers, you can also use the greater than and less than signs:"
+ "For numbers, you can also use the greater than and less than signs:\n"
]
},
{
@@ -372,7 +404,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df.query(\"col0 > 6\")"
+ "df.filter(pl.col(\"col0\") > 6)"
]
},
{
@@ -380,7 +412,7 @@
"id": "f5e03f63",
"metadata": {},
"source": [
- "In fact, there are lots of options that work with `query()`: as well as `>` (greater than), you can use `>=` (greater than or equal to), `<` (less than), `<=` (less than or equal to), `==` (equal to), and `!=` (not equal to). You can also use the commands `and` as well as `or` to combine multiple conditions. Here's an example of `and` from the `flights` data frame:"
+ "In fact, there are lots of options that work with `filter()`: as well as `>` (greater than), you can use `>=` (greater than or equal to), `<` (less than), `<=` (less than or equal to), `==` (equal to), and `!=` (not equal to). You can also use operators `&` as well as `|` to combine multiple conditions. Here's an example of `&` from the `flights` data frame:\n"
]
},
{
@@ -391,7 +423,7 @@
"outputs": [],
"source": [
"# Flights that departed on January 1\n",
- "flights.query(\"month == 1 and day == 1\")"
+ "flights.filter((pl.col(\"month\") == 1) & (pl.col(\"day\") <= 5))"
]
},
{
@@ -399,7 +431,7 @@
"id": "bd0af6fc",
"metadata": {},
"source": [
- "Note that equality is tested by `==` and *not* by `=`, because the latter is used for assignment."
+ "Note that equality is tested by `==` and _not_ by `=`, because the latter is used for assignment.\n"
]
},
{
@@ -409,7 +441,17 @@
"source": [
"### Re-arranging Rows\n",
"\n",
- "Again and again, you will want to re-order the rows of your data frame according to the values in a particular column. **pandas** makes this very easy via the `.sort_values()` function. It takes a data frame and a set of column names to sort by. If you provide more than one column name, each additional column will be used to break ties in the values of preceding columns. For example, the following code sorts by the departure time, which is spread over four columns."
+ "Again and again, you will want to re-order the rows of your data frame according to the values in a particular column. **polars** makes this very easy via the `.sort()` function. You can sort by single or multiple column names and also by expressions. If you provide more than one column name, each additional column will be used to break ties in the values of preceding columns. For example, the following code sorts by the departure time, which is spread over four columns.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "395c9c62",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "flights.sort(\"dep_time\")"
]
},
{
@@ -419,7 +461,11 @@
"metadata": {},
"outputs": [],
"source": [
- "flights.sort_values([\"year\", \"month\", \"day\", \"dep_time\"])"
+ "# Sort by multiple columns by passing a list of columns.\n",
+ "flights.sort([\"year\", \"month\", \"day\", \"dep_time\"])\n",
+ "\n",
+ "# Or use positional arguments to sort by multiple columns in the same way.\n",
+ "flights.sort(\"year\", \"month\", \"day\", \"dep_time\")"
]
},
{
@@ -427,8 +473,8 @@
"id": "39a6e9b1",
"metadata": {},
"source": [
- "You can use the keyword argument `ascending=False` to re-order by a column or columns in descending order.\n",
- "For example, this code shows the most delayed flights:"
+ "You can use the keyword argument `descending=True` to re-order by a column or columns in descending order.\n",
+ "For example, this code shows the most delayed flights:\n"
]
},
{
@@ -438,7 +484,17 @@
"metadata": {},
"outputs": [],
"source": [
- "flights.sort_values(\"dep_delay\", ascending=False)"
+ "flights.sort(\"dep_delay\", descending=True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "80bf3df7",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "flights.sort([\"dep_delay\", \"arr_delay\"], descending=[True, False])"
]
},
{
@@ -447,7 +503,7 @@
"metadata": {},
"source": [
"You can of course combine all of the above row manipulations to solve more complex problems.\n",
- "For example, we could look for the top three destinations of the flights that were most delayed on arrival that left on roughly on time:"
+ "For example, we could look for the top three destinations of the flights that were most delayed on arrival that left on roughly on time:\n"
]
},
{
@@ -458,9 +514,9 @@
"outputs": [],
"source": [
"(\n",
- " flights.query(\"dep_delay <= 10 and dep_delay >= -10\")\n",
- " .sort_values(\"arr_delay\", ascending=False)\n",
- " .iloc[[0, 1, 2]]\n",
+ " flights.filter((pl.col(\"dep_delay\") <= 10) & (pl.col(\"dep_delay\") >= -10))\n",
+ " .sort(\"arr_delay\", descending=True)\n",
+ " .head(3)\n",
")"
]
},
@@ -485,13 +541,13 @@
"\n",
" f. Were delayed by at least an hour, but made up over 30 minutes in flight\n",
"\n",
- "2. Sort `flights` to find the flights with longest departure delays.\n",
+ "2. Sort `flights` to find the flights with longest departure delays.\n",
"\n",
- "3. Sort `flights` to find the fastest flights\n",
+ "3. Sort `flights` to find the fastest flights\n",
"\n",
- "4. Which flights traveled the farthest?\n",
+ "4. Which flights traveled the farthest?\n",
"\n",
- "5. Does it matter what order you used `query()` and `sort_values()` in if you're using both? Why/why not? Think about the results and how much work the functions would have to do."
+ "5. Does it matter what order you used `filter()` and `sort()` in if you're using both? Why/why not? Think about the results and how much work the functions would have to do.\n"
]
},
{
@@ -505,7 +561,7 @@
"\n",
"::: {.callout-note}\n",
"Some **pandas** operations can apply either to columns or rows, depending on the syntax used. For example, accessing values by position can be achieved in the same way for rows and columns via `.iloc` where to access the ith row you would use `df.iloc[i]` and to access the jth column you would use `df.iloc[:, j]` where `:` stands in for 'any row'.\n",
- ":::"
+ ":::\n"
]
},
{
@@ -515,7 +571,7 @@
"source": [
"### Creating New Columns\n",
"\n",
- "Let's now move on to creating new columns, either using new information or from existing columns. Given a data frame, `df`, creating a new column with the same value repeated is as easy as using square brackets with a string (text enclosed by quotation marks) in."
+ "Let's now move on to creating new columns, either using new information or from existing columns. Given a data frame, `df`, creating a new column with the same value repeated is done by using `.with_columns()`, with an expression assigned to a column name. Here we use `pl.lit()` which returns an expression representing a literal value, 5 in our case.\n"
]
},
{
@@ -525,7 +581,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df[\"new_column0\"] = 5\n",
+ "df = df.with_columns(new_column0=pl.lit(5))\n",
"df"
]
},
@@ -534,7 +590,7 @@
"id": "55bc84a5",
"metadata": {},
"source": [
- "If we do the same operation again, but with a different right-hand side, it will overwrite what was already in that column. Let's see this with an example where we put different values in each position by assigning a list to the new column."
+ "If we do the same operation again, but this time assigning a **_Series_** to the same column, it will overwrite what was already in that column. A **_Series_** repesents a single column in a Polars DataFrame. Let's see this with an example where we put different values in each position by assigning a list to the new column.\n"
]
},
{
@@ -544,7 +600,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df[\"new_column0\"] = [0, 1, 2, 3, 4, 5]\n",
+ "df = df.with_columns(new_column0=pl.Series([0, 1, 2, 3, 4, 5]))\n",
"df"
]
},
@@ -557,7 +613,7 @@
"What happens if you try to use assignment where the right-hand side values are longer or shorter than the length of the data frame?\n",
":::\n",
"\n",
- "By passing a list within the square brackets, we can actually create more than one new column:"
+ "We can actually use `.with_columns` to create more than one new column:\n"
]
},
{
@@ -567,7 +623,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df[[\"new_column1\", \"new_column2\"]] = [5, 6]\n",
+ "df = df.with_columns(new_column1=pl.lit(5), new_column2=pl.lit(6))\n",
"df"
]
},
@@ -576,7 +632,7 @@
"id": "10792ddd",
"metadata": {},
"source": [
- "Very often, you will want to create a new column that is the result of an operation on existing columns. There are a couple of ways to do this. The 'stand-alone' method works in a similar way to what we've just seen except that we refer to the data frame on the right-hand side of the assignment statement too:"
+ "Very often, you will want to create a new column or modify a column that is the result of an operation on existing columns. There are a couple of ways to do this. The 'stand-alone' method works in a similar way to what we've just seen except that we refer to the data frame on the right-hand side of the assignment statement too:\n"
]
},
{
@@ -586,8 +642,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df[\"new_column3\"] = df[\"col0\"] - df[\"new_column0\"]\n",
- "df"
+ "df.with_columns(new_column2=pl.col(\"col0\") - pl.col(\"new_column0\"))"
]
},
{
@@ -595,7 +650,8 @@
"id": "03172fa9",
"metadata": {},
"source": [
- "The other way to do this involves an 'assign()' statement and is used when you wish to chain multiple steps together (like we saw earlier). These use a special syntax called a 'lambda' statement, which (here at least) just provides a way of specifying to **pandas** that we wish to perform the operation on every row. Below is an example using the flights data. You should note though that the word 'row' below is a dummy; you could replace it with any variable name (for example, `x`) but `row` makes what is happening a little bit clearer."
+ "We can use `.alias()` with an expression to assign column names, when creating new columns.\n",
+ "We can chain multiple expressions together with `.with_columns()`, which would create multiple new columns with the names assigned to `.alias()`.\n"
]
},
{
@@ -605,36 +661,18 @@
"metadata": {},
"outputs": [],
"source": [
- "(\n",
- " flights.assign(\n",
- " gain=lambda row: row[\"dep_delay\"] - row[\"arr_delay\"],\n",
- " speed=lambda row: row[\"distance\"] / row[\"air_time\"] * 60,\n",
- " )\n",
+ "flights.with_columns(\n",
+ " (pl.col(\"dep_delay\") - pl.col(\"arr_delay\")).alias(\"gain\"),\n",
+ " (pl.col(\"distance\") / pl.col(\"air_time\") * 60).alias(\"speed\"),\n",
")"
]
},
- {
- "cell_type": "markdown",
- "id": "c531df3e",
- "metadata": {},
- "source": [
- "::: {.callout-note}\n",
- "A lambda function is like any normal function in Python except that it has no name, and it tends to be contained in one line of code. A lambda function is made of an argument, a colon, and an expression, like the following lambda function that multiplies an input by three.\n",
- "\n",
- "```python\n",
- "lambda x: x*3\n",
- "```\n",
- "\n",
- ":::"
- ]
- },
{
"cell_type": "markdown",
"id": "82a97330",
"metadata": {},
"source": [
- "### Accessing Columns\n",
- "\n"
+ "### Accessing Columns\n"
]
},
{
@@ -642,7 +680,7 @@
"id": "7599db58",
"metadata": {},
"source": [
- "Just as with selecting rows, there are many options and ways to select the columns to operate on. The one with the simplest syntax is the name of the data frame followed by square brackets and the column name (as a string)"
+ "Just as with selecting rows, there are many options and ways to select the columns to operate on. The one with the simplest syntax is the name of the data frame followed by square brackets and the column name (as a string)\n"
]
},
{
@@ -660,7 +698,7 @@
"id": "63bca028",
"metadata": {},
"source": [
- "If you need to select *multiple* columns, you cannot just pass a string into `df[...]`; instead you need to pass an object that is iterable (and so have multiple items). The most straight forward way to select multiple columns is to pass a *list*. Remember, lists comes in square brackets so we're going to see something with repeated square brackets: one for accessing the data frame's innards and one for the list."
+ "If you need to select _multiple_ columns, you cannot just pass a string into `df[...]`; instead you need to pass an object that is iterable (and so have multiple items). The most straight forward way to select multiple columns is to pass a _list_. Remember, lists comes in square brackets so we're going to see something with repeated square brackets: one for accessing the data frame's innards and one for the list.\n"
]
},
{
@@ -675,46 +713,63 @@
},
{
"cell_type": "markdown",
- "id": "2b2a7be0",
+ "id": "a6fdfc17",
"metadata": {},
"source": [
- "If you want to access particular rows at the same time, use the `.loc` access function:"
+ "We can also use `.select()` on the data frame to select columns, passing a single string to select a single column or an iterable, like a _list_, _positional arguments_ or _keyword arguments_, to select multiple columns. **Using _keyword arguments_ renames the columns in the output**\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "eabfd313",
+ "id": "1bc0cd22",
"metadata": {},
"outputs": [],
"source": [
- "df.loc[[\"a\", \"b\"], [\"col0\", \"new_column0\", \"col2\"]]"
+ "# selecting a single column\n",
+ "df.select(\"col0\")\n",
+ "\n",
+ "# Using positional arguments to select multiple columns\n",
+ "df.select(\"col0\", \"new_column0\", \"col2\")\n",
+ "\n",
+ "# Using keyword arguments to rename columns in the output\n",
+ "df.select(col1=\"col0\", col2=\"new_column0\", col3=\"col2\")"
]
},
{
"cell_type": "markdown",
- "id": "c1b7db13",
+ "id": "a806be16",
"metadata": {},
"source": [
- "And, just as with rows, we can access columns by their position using `.iloc` (where `:` stands in for 'any row')."
+ "Expressions are also accepted\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "b6ae1605",
+ "id": "ed447fb7",
"metadata": {},
"outputs": [],
"source": [
- "df.iloc[:, [0, 1]]"
+ "df.select(pl.col(\"col0\"), pl.col(\"new_column0\") + 2, pl.col(\"col2\") * 2)"
]
},
{
"cell_type": "markdown",
- "id": "509dc236",
+ "id": "2b2a7be0",
+ "metadata": {},
+ "source": [
+ "If we want to access particular rows at the same time, we can chain `.filter()` or `.slice()` to the `.select()` function:\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "eabfd313",
"metadata": {},
+ "outputs": [],
"source": [
- "There are other ways to access multiple columns that make use of slicing but we’ll leave that topic for another time."
+ "df.select(\"col0\", \"new_column0\", \"col2\").slice(0, 2)"
]
},
{
@@ -722,46 +777,55 @@
"id": "17b928c8",
"metadata": {},
"source": [
- "Sometimes, you'll want to select columns based on the *type* of data that they hold. For this, **pandas** provides a function `.select_dtypes()`. Let's use this to select all columns with integers in the flights data."
+ "Sometimes, we'll want to select columns based on the _type_ of data that they hold. For this, we can call **polars** data types with an expression inside `.select()`. Let's use this to select all columns with integers in the flights data.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "62f578d1",
+ "id": "aed67406",
"metadata": {},
"outputs": [],
"source": [
- "flights.select_dtypes(\"int\")"
+ "flights.select(pl.col(pl.Int64))"
]
},
{
"cell_type": "markdown",
- "id": "9aec778c",
+ "id": "8cb930af",
"metadata": {},
"source": [
- "There are other occasions when you'd like to select columns based on criteria such as patterns in the *name* of the column. Because Python has very good support for text, this is very possible but doesn't tend to be so built-in to **pandas** functions. The trick is to generate a list of column names that you want from the pattern you're interested in.\n",
- "\n",
- "Let's see a couple of examples. First, let's get all columns in our `df` data frame that begin with `\"new_...\"`. We'll generate a list of true and false values reflecting if each of the columns begins with \"new\" and then we'll pass those true and false values to `.loc`, which will only give columns for which the result was `True`. To show what's going on, we'll break it into two steps:"
+ "**polars** also provides a `selectors` module that we can use to select columns based on both data types and criteria such as patterns in the name of the column.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "5aaae8bd",
+ "id": "62f578d1",
"metadata": {},
"outputs": [],
"source": [
- "print(\"The list of columns:\")\n",
- "print(df.columns)\n",
- "print(\"\\n\")\n",
+ "import polars.selectors as S\n",
"\n",
- "print(\"The list of true and false values:\")\n",
- "print(df.columns.str.startswith(\"new\"))\n",
- "print(\"\\n\")\n",
+ "# Select all integer columns\n",
+ "flights.select(S.integer())\n",
+ "\n",
+ "# Exclude string columns\n",
+ "flights.select(S.exclude(S.string()))"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "id": "d4e486db",
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Select columns that contain \"delay\" in their name\n",
+ "flights.select(S.contains(\"delay\"))\n",
"\n",
- "print(\"The selection from the data frame:\")\n",
- "df.loc[:, df.columns.str.startswith(\"new\")]"
+ "# Select columns that start with \"arr\"\n",
+ "flights.select(S.starts_with(\"arr\"))"
]
},
{
@@ -769,7 +833,7 @@
"id": "b514cbf4",
"metadata": {},
"source": [
- "As well as `startswith()`, there are other commands like `endswith()`, `contains()`, `isnumeric()`, and `islower()`."
+ "Other `selectors` commands include `ends_with()`, `by_index()`, `first()`, `last()`, `duration()`, `numeric()`, `boolean()` and more\n"
]
},
{
@@ -779,7 +843,7 @@
"source": [
"### Renaming Columns\n",
"\n",
- "There are three easy ways to rename columns, depending on what the context is. The first is to use the dedicated `rename()` function with an object called a dictionary. Dictionaries in Python consist of curly brackets with comma separated pairs of values where the first values maps into the second value. An example of a dictionary would be `{'old_col1': 'new_col1', 'old_col2': 'new_col2'}`. Let's see this in practice (but note that we are not 'saving' the resulting data frame, just showing it—to save it, you'd need to add `df = ` to the left-hand side of the code below)."
+ "We use the dedicated `rename()` function with a mapping, such as a dictionary or a lambda function. Dictionaries in Python consist of curly brackets with comma separated pairs of values where the first values maps into the second value. An example of a dictionary would be `{'old_col1': 'new_col1', 'old_col2': 'new_col2'}`. Let's see this in practice (but note that we are not 'saving' the resulting data frame, just showing it—to save it, you'd need to add `df = ` to the left-hand side of the code below).\n"
]
},
{
@@ -789,7 +853,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df.rename(columns={\"col3\": \"letters\", \"col4\": \"names\", \"col6\": \"fruit\"})"
+ "df.rename({\"col3\": \"letters\", \"col4\": \"names\", \"col6\": \"fruit\"})"
]
},
{
@@ -797,7 +861,7 @@
"id": "0a673852",
"metadata": {},
"source": [
- "The second method is for when you want to rename all of the columns. For that you simply set `df.columns` equal to the new set of columns that you'd like to have. For example, we might want to capitalise the first letter of each column using `str.capitalize()` and assign that to `df.columns`."
+ "Using a lambda function, maps each column name as its argument, which you can then perform an operation on.\n"
]
},
{
@@ -807,26 +871,16 @@
"metadata": {},
"outputs": [],
"source": [
- "df.columns = df.columns.str.capitalize()\n",
- "df"
+ "df.rename(lambda column_name: column_name.upper())"
]
},
{
"cell_type": "markdown",
- "id": "7a8b9660",
- "metadata": {},
- "source": [
- "Finally, we might be interested in just replacing specific parts of column names. In this case, we can use `.str.replace()`. As an example, let's add the word `\"Original\"` ahead of the original columns:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "id": "3dd7606b",
+ "id": "aa80d44b",
"metadata": {},
- "outputs": [],
"source": [
- "df.columns.str.replace(\"Col\", \"Original_column\")"
+ "::: {.callout-tip}\n",
+ "A _lambda function_ is a small, anonymous function in Python that performs a single operation. It's a shorthand way to create a function without using `def`.\n"
]
},
{
@@ -834,7 +888,7 @@
"id": "09632b99",
"metadata": {},
"source": [
- "### Re-ordering Columns"
+ "### Re-ordering Columns\n"
]
},
{
@@ -844,9 +898,9 @@
"source": [
"By default, new columns are added to the right-hand side of the data frame. But you may have reasons to want the columns to appear in a particular order, or perhaps you'd just find it more convenient to have new columns on the left-hand side when there are many columns in a data frame (which happens a lot).\n",
"\n",
- "The simplest way to re-order (all) columns is to create a new list of their names with them in the order that you'd like them: but be careful you don't forget any columns that you'd like to keep! \n",
+ "The simplest way to re-order (all) columns is to create a new list of their names with them in the order that you'd like them: but be careful you don't forget any columns that you'd like to keep!\n",
"\n",
- "Let's see an example with a fresh version of the fake data from earlier. We'll put all of the odd-numbered columns first, in descending order, then the even similarly."
+ "Let's see an example with a fresh version of the fake data from earlier. We'll put all of the odd-numbered columns first, in descending order, then the even similarly using `.select()`.\n"
]
},
{
@@ -856,11 +910,8 @@
"metadata": {},
"outputs": [],
"source": [
- "df = pd.DataFrame(\n",
- " data=np.reshape(range(36), (6, 6)),\n",
- " index=[\"a\", \"b\", \"c\", \"d\", \"e\", \"f\"],\n",
- " columns=[\"col\" + str(i) for i in range(6)],\n",
- " dtype=float,\n",
+ "df = pl.DataFrame(\n",
+ " data=np.reshape(range(36), (6, 6)), schema=[\"col\" + str(i) for i in range(6)]\n",
")\n",
"df"
]
@@ -868,11 +919,11 @@
{
"cell_type": "code",
"execution_count": null,
- "id": "b9a409ac",
+ "id": "3c2029cc",
"metadata": {},
"outputs": [],
"source": [
- "df = df[[\"col5\", \"col3\", \"col1\", \"col4\", \"col2\", \"col0\"]]\n",
+ "df = df.select([\"col5\", \"col3\", \"col1\", \"col4\", \"col2\", \"col0\"])\n",
"df"
]
},
@@ -881,7 +932,7 @@
"id": "dd91d87a",
"metadata": {},
"source": [
- "Of course, this is quite tedious if you have lots of columns! There are methods that can help make this easier depending on your context. Perhaps you'd just liked to sort the columns in order? This can be achieved by combining `sorted()` and the `reindex()` command (which works for rows or columns) with `axis=1`, which means the second axis (i.e. columns)."
+ "Of course, this is quite tedious if you have lots of columns! There are methods that can help make this easier depending on your context. Perhaps you'd just liked to sort the columns in order? This can be achieved by combining `sorted()` and the `.select()` function, passing the DataFrame `.columns`.\n"
]
},
{
@@ -891,47 +942,11 @@
"metadata": {},
"outputs": [],
"source": [
- "df.reindex(sorted(df.columns), axis=1)"
- ]
- },
- {
- "cell_type": "markdown",
- "id": "28e49605",
- "metadata": {},
- "source": [
- "## Review of How to Access Rows, Columns, and Values\n",
- "\n",
- "With all of these different ways to access values in data frames, it can get confusing. These are the different ways to get the first column of a data frame (when that first column is called `column` and the data frame is `df`):\n",
- "\n",
- "- `df.column`\n",
- "- `df[\"column\"]`\n",
- "- `df.loc[:, \"column\"]`\n",
- "- `df.iloc[:, 0]`\n",
- "\n",
- "Note that `:` means 'give me everything'! The ways to access rows are similar (here assuming the first row is called `row`):\n",
- "\n",
- "- `df.loc[\"row\", :]`\n",
- "- `df.iloc[0, :]`\n",
- "\n",
- "And to access the first value (ie the value in first row, first column):\n",
- "\n",
- "- `df.column[0]`\n",
- "- `df[\"column\"][0]`\n",
- "- `df.iloc[0, 0]`\n",
- "- `df.loc[\"row\", \"column\"]`\n",
- "\n",
- "In the above examples, square brackets are instructions about *where* to grab bits from the data frame. They are a bit like an address system for values within a data frame. Square brackets *also* denote lists though. So if you want to select *multiple* columns or rows, you might see syntax like this:\n",
- "\n",
- "`df.loc[[\"row0\", \"row1\"], [\"column0\", \"column2\"]]`\n",
- "\n",
- "which picks out two rows and two columns via the lists `[\"row0\", \"row1\"]` and `[\"column0\", \"column2\"]`. Because there are lists alongside the usual system of selecting values, there are two sets of square brackets.\n",
- "\n",
- "::: {.callout-tip title=\"Tip\"}\n",
+ "# Alphabetical order\n",
+ "df.select(sorted(df.columns))\n",
"\n",
- "If you only want to remember one syntax for accessing rows and columns by name, use the pattern `df.loc[[\"row0\", \"row1\", ...], [\"col0\", \"col1\", ...]]`. This also works with a single row or a single column (or both).\n",
- "\n",
- "If you only want to remember one syntax for accessing rows and columns by position, use the pattern `df.iloc[[0, 1, ...], [0, 1, ...]]`. This also works with a single row or a single column (or both).\n",
- ":::\n"
+ "# Reverse alphabetical order\n",
+ "df.select(sorted(df.columns, reverse=True))"
]
},
{
@@ -941,29 +956,13 @@
"source": [
"### Column and Row Exercises\n",
"\n",
- "1. Compare `air_time` with `arr_time - dep_time`. What do you expect to see? What do you see What do you need to do to fix it?\n",
+ "1. Compare `air_time` with `arr_time - dep_time`. What do you expect to see? What do you see? What do you need to do to fix it?\n",
"\n",
"2. Compare `dep_time`, `sched_dep_time`, and `dep_delay`. How would you expect those three numbers to be related?\n",
"\n",
"3. Brainstorm as many ways as possible to select `dep_time`, `dep_delay`, `arr_time`, and `arr_delay` from `flights`.\n",
"\n",
- "4. What happens if you include the name of a row or column multiple times when trying to select them?\n",
- "\n",
- "5. What does the `.isin()` function do in the following?\n",
- "\n",
- " ```python\n",
- " flights.columns.isin([\"year\", \"month\", \"day\", \"dep_delay\", \"arr_delay\"])\n",
- " ```\n",
- "\n",
- "6. Does the result of running the following code surprise you?\n",
- " How do functions like `str.contains` deal with case by default?\n",
- " How can you change that default?\n",
- "\n",
- " ```python\n",
- " flights.loc[:, flights.columns.str.contains(\"TIME\")]\n",
- " ```\n",
- "\n",
- " (Hint: you can use help even on functions that apply to data frames, eg use `help(flights.columns.str.contains)`)"
+ "4. What happens if you include the name of a row or column multiple times when trying to select them?\n"
]
},
{
@@ -971,9 +970,9 @@
"id": "a3c837e4",
"metadata": {},
"source": [
- "## Grouping, changing the index, and applying summary statistics\n",
+ "## Grouping and applying summary statistics\n",
"\n",
- "So far you've learned about working with rows and columns. **pandas** gets even more powerful when you add in the ability to work with groups. Creating groups will often also mean a change of index. And because groups tend to imply an aggregation or pooling of data, they often go hand-in-hand with the application of a summary statistic.\n",
+ "So far you've learned about working with rows and columns. **polars** gets even more powerful when you add in the ability to work with groups. And because groups tend to imply an aggregation or pooling of data, they go hand-in-glove with the application of a summary statistic.\n",
"\n",
"The diagram below gives a sense of how these operations can proceed together. Note that the 'split' operation is achieved through grouping, while apply produces summary statistics. At the end, you get a data frame with a new index (one entry per group) in what is shown as the 'combine' step.\n",
"\n",
@@ -981,9 +980,7 @@
"\n",
"### Grouping and Aggregating\n",
"\n",
- "Let's take a look at creating a group using the `.groupby()` function followed by selecting a column and applying a summary statistic via an aggregation. Note that *aggregation*, via `.agg()`, always produces a new index because we have collapsed information down to the group-level (and the new index is made of those levels).\n",
- "\n",
- "The key point to remember is: use `.agg()` with `.groupby()` when you want your groups to become the new index."
+ "Let's take a look at creating a group using the `.group_by()` function, then followed by the `.agg()` function for aggregation, selecting a column and applying a summary statistic via an aggregation.\n"
]
},
{
@@ -993,7 +990,7 @@
"metadata": {},
"outputs": [],
"source": [
- "(flights.groupby(\"month\")[[\"dep_delay\"]].mean())"
+ "flights.group_by(\"month\").agg(pl.col(\"dep_delay\").mean())"
]
},
{
@@ -1001,19 +998,9 @@
"id": "b003ea0d",
"metadata": {},
"source": [
- "This now represents the mean departure delay by month. Notice that our index has changed! We now have month where we original had an index that was just the row number. The index plays an important role in grouping operations because it keeps track of the groups you have in the rest of your data frame.\n",
+ "This now represents the mean departure delay by month. The mechanics happenning here is that the DataFrame is grouped by each unique item in the _\"month\"_ column and then a mean summary statistic is derived on the _\"dep_delay\"_ column from each group.\n",
"\n",
- "Often, you might want to do multiple summary operations in one go. The most comprehensive syntax for this is via `.agg()`. We can reproduce what we did above using `.agg()`:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "id": "aaaca266",
- "metadata": {},
- "outputs": [],
- "source": [
- "(flights.groupby(\"month\")[[\"dep_delay\"]].agg(\"mean\"))"
+ "Other summary statistics can be derived from aggregations. Some common options are in the table below:\n"
]
},
{
@@ -1021,21 +1008,20 @@
"id": "520299c7",
"metadata": {},
"source": [
- "where you pass in whatever aggregation you want. Some common options are in the table below:\n",
- "\n",
- "| Aggregation | Description |\n",
- "| ----------- | ----------- |\n",
- "| `count()` | Number of items |\n",
- "| `first()`, `last()` | \tFirst and last item |\n",
- "| `mean()`, `median()` |\tMean and median |\n",
- "| `min()`, `max()` |\tMinimum and maximum |\n",
- "| `std()`, `var()` |\tStandard deviation and variance |\n",
- "| `mad()` |\tMean absolute deviation |\n",
- "| `prod()` |\tProduct of all items |\n",
- "| `sum()`\t| Sum of all items |\n",
- "| `value_counts()` | Counts of unique values |\n",
+ "| Aggregation | Description |\n",
+ "| -------------------- | ------------------------------- |\n",
+ "| `count()` | Number of non-null items |\n",
+ "| `len()` | Number of all items |\n",
+ "| `first()`, `last()` | First and last item |\n",
+ "| `mean()`, `median()` | Mean and median |\n",
+ "| `min()`, `max()` | Minimum and maximum |\n",
+ "| `std()`, `var()` | Standard deviation and variance |\n",
+ "| `mad()` | Mean absolute deviation |\n",
+ "| `product()` | Product of all items |\n",
+ "| `sum()` | Sum of all items |\n",
+ "| `value_counts()` | Counts of unique values |\n",
"\n",
- "For doing multiple aggregations on multiple columns with new names for the output variables, the syntax becomes"
+ "For doing multiple aggregations on multiple columns with new names for the output variables, the syntax becomes\n"
]
},
{
@@ -1045,11 +1031,10 @@
"metadata": {},
"outputs": [],
"source": [
- "(\n",
- " flights.groupby([\"month\"]).agg(\n",
- " mean_delay=(\"dep_delay\", \"mean\"),\n",
- " count_flights=(\"dep_delay\", \"count\"),\n",
- " )\n",
+ "# Multiple aggregations using polars' syntactic sugar (shorthand) for mean and count\n",
+ "flights.group_by(\"month\").agg(\n",
+ " mean_delay=pl.mean(\"dep_delay\"),\n",
+ " count_flights=pl.count(\"dep_delay\"),\n",
")"
]
},
@@ -1058,7 +1043,7 @@
"id": "c331e813",
"metadata": {},
"source": [
- "Means and counts can get you a surprisingly long way in data science!"
+ "Means and counts can get you a surprisingly long way in data science!\n"
]
},
{
@@ -1068,7 +1053,7 @@
"source": [
"### Grouping by multiple variables\n",
"\n",
- "This is as simple as passing `.groupby()` a list representing multiple columns instead of a string representing a single column."
+ "This is as simple as passing `.group_by()` a list or multiple strings representing columns instead of a single string representing a single column.\n"
]
},
{
@@ -1078,95 +1063,13 @@
"metadata": {},
"outputs": [],
"source": [
- "month_year_delay = flights.groupby([\"month\", \"year\"]).agg(\n",
- " mean_delay=(\"dep_delay\", \"mean\"),\n",
- " count_flights=(\"dep_delay\", \"count\"),\n",
+ "month_year_delay = flights.group_by(\"month\", \"year\").agg(\n",
+ " mean_delay=pl.mean(\"dep_delay\"),\n",
+ " count_flights=pl.count(\"dep_delay\"),\n",
")\n",
"month_year_delay"
]
},
- {
- "cell_type": "markdown",
- "id": "22e89b7e",
- "metadata": {},
- "source": [
- "You might have noticed that this time we have a multi-index (that is, an index with more than one column). That's because we asked for something with multiple groups, and the index tracks what's going on within each group: so we need more than one dimension of index to do this efficiently.\n",
- "\n",
- "If you ever want to go back to an index that is just the position, try `reset_index()`"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "id": "33712dc2",
- "metadata": {},
- "outputs": [],
- "source": [
- "month_year_delay.reset_index()"
- ]
- },
- {
- "cell_type": "markdown",
- "id": "96c8416a",
- "metadata": {},
- "source": [
- "Perhaps you only want to remove one layer of the index though. This can be achieved by passing the position of the index you'd like to remove: for example, to only change the year index to a column, we would use: "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "id": "bcbef8f7",
- "metadata": {},
- "outputs": [],
- "source": [
- "month_year_delay.reset_index(1)"
- ]
- },
- {
- "cell_type": "markdown",
- "id": "8ab554b2",
- "metadata": {},
- "source": [
- "Finally, you can do more complicated re-arrangements of the index with an operation called `unstack`, which pivots the chosen index variable to be a column variable instead (introducing a multi column level structure). It's usually best to avoid this."
- ]
- },
- {
- "cell_type": "markdown",
- "id": "b3e1ea60",
- "metadata": {},
- "source": [
- "### Grouping and Transforming\n",
- "\n",
- "You may not always want to change the index to reflect new groups when performing computations at the group level.\n",
- "\n",
- "The key point to remember is: use `.transform()` with `.groupby()` when you want to perform computations on your groups but you want to go back to the original index.\n",
- "\n",
- "Let's say we wanted to express the arrival delay, `\"arr_del\"`, of each flight as a fraction of the worst arrival delay in each month."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "id": "f96fcf41",
- "metadata": {},
- "outputs": [],
- "source": [
- "flights[\"max_delay_month\"] = flights.groupby(\"month\")[\"arr_delay\"].transform(\"max\")\n",
- "flights[\"delay_frac_of_max\"] = flights[\"arr_delay\"] / flights[\"max_delay_month\"]\n",
- "flights[\n",
- " [\"year\", \"month\", \"day\", \"arr_delay\", \"max_delay_month\", \"delay_frac_of_max\"]\n",
- "].head()"
- ]
- },
- {
- "cell_type": "markdown",
- "id": "3880ece9",
- "metadata": {},
- "source": [
- "Note that the first few entries of `\"max_delay_month\"` are all the same because the month is the same for those entries, but the delay fraction changes with each row."
- ]
- },
{
"cell_type": "markdown",
"id": "1954cb3b",
@@ -1174,18 +1077,15 @@
"source": [
"### Groupby Exercises\n",
"\n",
- "1. Which carrier has the worst delays? Challenge: can you disentangle the effects of bad airports vs. bad carriers? Why/why not? (Hint: think about `flights.groupby([\"carrier\", \"dest\"]).count()`)\n",
+ "1. Which carrier has the worst delays? Challenge: can you disentangle the effects of bad airports vs. bad carriers? Why/why not? (Hint: think about `flights.group_by([\"carrier\", \"dest\"])`)\n",
"\n",
"2. Find the most delayed flight to each destination.\n",
"\n",
- "3. How do delays vary over the course of the day?"
+ "3. How do delays vary over the course of the day?\n"
]
}
],
"metadata": {
- "interpreter": {
- "hash": "9d7534ecd9fbc7d385378f8400cf4d6cb9c6175408a574f1c99c5269f08771cc"
- },
"jupytext": {
"cell_metadata_filter": "-all",
"encoding": "# -*- coding: utf-8 -*-",
@@ -1193,7 +1093,7 @@
"main_language": "python"
},
"kernelspec": {
- "display_name": "Python 3 (ipykernel)",
+ "display_name": "python4ds",
"language": "python",
"name": "python3"
},
diff --git a/data-visualise.ipynb b/data-visualise.ipynb
index 0335c48..488a1db 100644
--- a/data-visualise.ipynb
+++ b/data-visualise.ipynb
@@ -17,7 +17,7 @@
"\n",
"However, we'll get further faster by learning one system and applying it in many places—and the beauty of declarative plotting is that it covers lots of standard charts simply and well. **letsplot** implements the so-called **grammar of graphics**, a coherent declarative system for describing and building graphs.\n",
"\n",
- "We will start by creating a simple scatterplot and use that to introduce aesthetic mappings and geometric objects—the fundamental building blocks of **letsplot**. We will then walk you through visualising distributions of single variables as well as visualising relationships between two or more variables. We’ll finish off with saving your plots and troubleshooting tips. "
+ "We will start by creating a simple scatterplot and use that to introduce aesthetic mappings and geometric objects—the fundamental building blocks of **letsplot**. We will then walk you through visualising distributions of single variables as well as visualising relationships between two or more variables. We’ll finish off with saving your plots and troubleshooting tips.\n"
]
},
{
@@ -27,7 +27,7 @@
"source": [
"### Prerequisites\n",
"\n",
- "You will need to install the **letsplot** package for this chapter. To do this, open up the command line of your computer, type in `uv add lets-plot`, and hit enter."
+ "You will need to install the **letsplot** package for this chapter. To do this, open up the command line of your computer, type in `uv add lets-plot`, and hit enter.\n"
]
},
{
@@ -39,7 +39,7 @@
"The command line can be opened within Visual Studio Code and Codespaces by going to View -> Terminal.\n",
":::\n",
"\n",
- "Note that you only need to install a package once in each Python environment."
+ "Note that you only need to install a package once in each Python environment.\n"
]
},
{
@@ -47,9 +47,9 @@
"id": "e0ad70c8",
"metadata": {},
"source": [
- "We'll also need to have the **pandas** package installed—this package, which we'll be seeing a lot of, is for data. You can similarly install it by running `uv add pandas` on the command line.\n",
+ "We'll also need to have the **polars** package installed—this package, which we'll be seeing a lot of, is for data. You can similarly install it by running `uv add polars` on the command line.\n",
"\n",
- "Finally, we'll also need some data (you can't science without data). We'll be using the Palmer penguins dataset. Unusually, this can also be installed as a package—normally you would load data from a file, but these data are so popular for tutorials they've found their way into an installable package. Run `uv add palmerpenguins` to get these data."
+ "Finally, we'll also need some data (you can't science without data). We'll be using the Palmer penguins dataset. Unusually, this can also be installed as a package—normally you would load data from a file, but these data are so popular for tutorials they've found their way into an installable package. Run `uv add palmerpenguins` to get these data.\n"
]
},
{
@@ -57,7 +57,7 @@
"id": "8852373a",
"metadata": {},
"source": [
- "Our next task is to load these into our Python session, either in a Python notebook cell within a Jupyter Notebook, by writing it in a script that we then send to the interactive window, or by typing it directly into the interactive window and hitting shift and enter. Here's the code:"
+ "Our next task is to load these into our Python session, either in a Python notebook cell within a Marimo or Jupyter Notebook, by writing it in a script that we then send to the interactive window, or by typing it directly into the interactive window and hitting shift and enter. Here's the code:\n"
]
},
{
@@ -67,6 +67,7 @@
"metadata": {},
"outputs": [],
"source": [
+ "import polars as pl\n",
"from lets_plot import *\n",
"from palmerpenguins import load_penguins\n",
"\n",
@@ -78,7 +79,7 @@
"id": "4443f4dd",
"metadata": {},
"source": [
- "These lines import parts of the **pandas** and **palmerpenguins** packages, then import all (`*`) of the functions of the **letsplot** package. The final line allows charts to display in HTML."
+ "These lines import parts of the **palmerpenguins** package, then import all (`*`) of the functions of the **letsplot** package. The final line allows charts to display in HTML.\n"
]
},
{
@@ -88,7 +89,7 @@
"source": [
"## First Steps\n",
"\n",
- "Do penguins with longer flippers weigh more or less than penguins with shorter flippers? You probably already have an answer, but try to make your answer precise. What does the relationship between flipper length and body mass look like? Is it positive? Negative? Linear? Nonlinear? Does the relationship vary by the species of the penguin? How about by the island where the penguin lives? Let’s create visualisations that we can use to answer these questions."
+ "Do penguins with longer flippers weigh more or less than penguins with shorter flippers? You probably already have an answer, but try to make your answer precise. What does the relationship between flipper length and body mass look like? Is it positive? Negative? Linear? Nonlinear? Does the relationship vary by the species of the penguin? How about by the island where the penguin lives? Let’s create visualisations that we can use to answer these questions.\n"
]
},
{
@@ -102,21 +103,21 @@
"\n",
"To make the discussion easier, let's define some terms:\n",
"\n",
- "- A **variable** is a quantity, quality, or property that you can measure.\n",
+ "- A **variable** is a quantity, quality, or property that you can measure.\n",
"\n",
- "- A **value** is the state of a variable when you measure it.\n",
- " The value of a variable may change from measurement to measurement.\n",
+ "- A **value** is the state of a variable when you measure it.\n",
+ " The value of a variable may change from measurement to measurement.\n",
"\n",
- "- An **observation** is a set of measurements made under similar conditions (you usually make all of the measurements in an observation at the same time and on the same object).\n",
- " An observation will contain several values, each associated with a different variable.\n",
- " We'll sometimes refer to an observation as a data point.\n",
+ "- An **observation** is a set of measurements made under similar conditions (you usually make all of the measurements in an observation at the same time and on the same object).\n",
+ " An observation will contain several values, each associated with a different variable.\n",
+ " We'll sometimes refer to an observation as a data point.\n",
"\n",
- "- **Tabular data** is a set of values, each associated with a variable and an observation.\n",
- " Tabular data is *tidy* if each value is placed in its own \"cell\", each variable in its own column, and each observation in its own row.\n",
+ "- **Tabular data** is a set of values, each associated with a variable and an observation.\n",
+ " Tabular data is _tidy_ if each value is placed in its own \"cell\", each variable in its own column, and each observation in its own row.\n",
"\n",
"In this context, a variable refers to an attribute of all the penguins, and an observation refers to all the attributes of a single penguin.\n",
"\n",
- "Type the name of the data frame in the interactive window and Python will print a preview of its contents."
+ "Type the name of the data frame in the interactive window and Python will print a preview of its contents.\n"
]
},
{
@@ -126,7 +127,7 @@
"metadata": {},
"outputs": [],
"source": [
- "penguins = load_penguins()\n",
+ "penguins = pl.from_pandas(load_penguins())\n",
"penguins"
]
},
@@ -135,7 +136,7 @@
"id": "cc310b4f",
"metadata": {},
"source": [
- "For an alternative view, where you can see the first few observations of each variable, use `penguins.head()`."
+ "For an alternative view, where you can see the first few observations of each variable, use `penguins.head()`.\n"
]
},
{
@@ -171,7 +172,7 @@
"source": [
"### Ultimate Goal\n",
"\n",
- "Our ultimate goal in this chapter is to recreate the following visualisation displaying the relationship between flipper lengths and body masses of these penguins, taking into consideration the species of the penguin."
+ "Our ultimate goal in this chapter is to recreate the following visualisation displaying the relationship between flipper lengths and body masses of these penguins, taking into consideration the species of the penguin.\n"
]
},
{
@@ -241,7 +242,7 @@
"For example, bar charts use bar geoms (`geom_bar()`), line charts use line geoms (`geom_line()`), boxplots use boxplot geoms (`geom_boxplot()`), scatterplots use point geoms (`geom_point()`), and so on.\n",
"\n",
"The function `geom_point()` adds a layer of points to your plot, which creates a scatterplot.\n",
- "**letsplot** comes with many geom functions that each adds a different type of layer to a plot."
+ "**letsplot** comes with many geom functions that each adds a different type of layer to a plot.\n"
]
},
{
@@ -266,7 +267,7 @@
"It doesn't yet match our \"ultimate goal\" plot, but using this plot we can start answering the question that motivated our exploration: \"What does the relationship between flipper length and body mass look like?\" The relationship appears to be positive (as flipper length increases, so does body mass), fairly linear (the points are clustered around a line instead of a curve), and moderately strong (there isn't too much scatter around such a line).\n",
"Penguins with longer flippers are generally larger in terms of their body mass.\n",
"\n",
- "It's a good point to flag that although we have plotted everything in the `penguins` data frame, there were a couple of rows with undefined values—and of course these cannot be plotted."
+ "It's a good point to flag that although we have plotted everything in the `penguins` data frame, there were a couple of rows with undefined values—and of course these cannot be plotted.\n"
]
},
{
@@ -285,7 +286,7 @@
"If you guessed \"in the aesthetic mapping, inside of `aes()`\", you're already getting the hang of creating data visualisations with **letsplot**!\n",
"And if not, don't worry.\n",
"\n",
- "Throughout the book you will make many more plots and have many more opportunities to check your intuition as you make them."
+ "Throughout the book you will make many more plots and have many more opportunities to check your intuition as you make them.\n"
]
},
{
@@ -319,7 +320,7 @@
"\n",
"Since this is a new geometric object representing our data, we will add a new geom as a layer on top of our point geom: `geom_smooth()`.\n",
"\n",
- "And we will specify that we want to draw the line of best fit based on a `l`inear `m`odel with `method = \"lm\"`."
+ "And we will specify that we want to draw the line of best fit based on a `l`inear `m`odel with `method = \"lm\"`.\n"
]
},
{
@@ -346,9 +347,9 @@
"source": [
"We have successfully added lines, but this plot doesn't look like the plot from earlier as that only had one line for the entire dataset as opposed to separate lines for each of the penguin species.\n",
"\n",
- "When aesthetic mappings are defined in `ggplot()`, at the *global* level, they're passed down to each of the subsequent geom layers of the plot.\n",
+ "When aesthetic mappings are defined in `ggplot()`, at the _global_ level, they're passed down to each of the subsequent geom layers of the plot.\n",
"\n",
- "However, each geom function in **letplot** can also take a `mapping` argument, which allows for aesthetic mappings at the *local* level that are added to those inherited from the global level.\n",
+ "However, each geom function in **letplot** can also take a `mapping` argument, which allows for aesthetic mappings at the _local_ level that are added to those inherited from the global level.\n",
"\n",
"Since we want points to be colored based on species but don't want the lines to be separated out for them, we should specify `color = species` for `geom_point()` only: therefore we take it out of the global `aes()` and just add it to `geom_point()`.\n"
]
@@ -376,7 +377,7 @@
"\n",
"We still need to use different shapes for each species of penguins and improve labels.\n",
"\n",
- "It's generally not a good idea to represent information using only colors on a plot, as people perceive colors differently due to color blindness or other color vision differences. Therefore, in addition to color, we can also map `species` to the `shape` aesthetic."
+ "It's generally not a good idea to represent information using only colors on a plot, as people perceive colors differently due to color blindness or other color vision differences. Therefore, in addition to color, we can also map `species` to the `shape` aesthetic.\n"
]
},
{
@@ -400,7 +401,7 @@
"source": [
"Note that the legend is automatically updated to reflect the different shapes of the points as well.\n",
"\n",
- "And finally, we can improve the labels of our plot using the `labs()` function in a new layer. Some of the arguments to `labs()` might be self explanatory: `title` adds a title and `subtitle` adds a subtitle to the plot. Other arguments match the aesthetic mappings, `x` is the x-axis label, `y` is the y-axis label, and `color` and `shape` define the label for the legend."
+ "And finally, we can improve the labels of our plot using the `labs()` function in a new layer. Some of the arguments to `labs()` might be self explanatory: `title` adds a title and `subtitle` adds a subtitle to the plot. Other arguments match the aesthetic mappings, `x` is the x-axis label, `y` is the y-axis label, and `color` and `shape` define the label for the legend.\n"
]
},
{
@@ -430,7 +431,7 @@
"id": "cdc33b33",
"metadata": {},
"source": [
- "We finally have a plot that perfectly matches our \"ultimate goal\"!"
+ "We finally have a plot that perfectly matches our \"ultimate goal\"!\n"
]
},
{
@@ -456,7 +457,7 @@
"5. Why does the following give an error and how would you fix it?\n",
"\n",
" ```python\n",
- " (ggplot(data = penguins) + \n",
+ " (ggplot(data = penguins) +\n",
" geom_point())\n",
" ```\n",
"\n",
@@ -464,7 +465,7 @@
"\n",
"7. Recreate the following visualisation.\n",
" What aesthetic should `bill_depth_mm` be mapped to?\n",
- " And should it be mapped at the global level or at the geom level?"
+ " And should it be mapped at the global level or at the geom level?\n"
]
},
{
@@ -490,7 +491,6 @@
"id": "986fdc29",
"metadata": {},
"source": [
- "\n",
"8. Run this code in your head and predict what the output will look like.\n",
" Then, run the code in Python and check your predictions.\n",
"\n",
@@ -518,6 +518,7 @@
" geom_smooth()\n",
" )\n",
" ```\n",
+ "\n",
" ```python\n",
" (ggplot() +\n",
" geom_point(\n",
@@ -529,7 +530,7 @@
" mapping = aes(x = \"flipper_length_mm\", y = \"body_mass_g\")\n",
" )\n",
" )\n",
- " ```"
+ " ```\n"
]
},
{
@@ -549,7 +550,7 @@
" mapping = aes(x = \"flipper_length_mm\", y = \"body_mass_g\")\n",
") +\n",
" geom_point())\n",
- "```"
+ "```\n"
]
},
{
@@ -565,10 +566,10 @@
"\n",
"```python\n",
"(\n",
- " ggplot(penguins, aes(x = \"flipper_length_mm\", y = \"body_mass_g\")) + \n",
+ " ggplot(penguins, aes(x = \"flipper_length_mm\", y = \"body_mass_g\")) +\n",
" geom_point()\n",
")\n",
- "```"
+ "```\n"
]
},
{
@@ -602,9 +603,9 @@
"id": "699f42eb",
"metadata": {},
"source": [
- "You may have seen earlier that the *data type* of the `\"species\"` column is string. Ideally, we want it to be categorical, so that there's no confusion about the fact that we're dealing with a finite number of mutually exclusive groups here. Another advantage is that it allows plotting tools to realise what kind of data it is working with.\n",
+ "You may have seen earlier that the _data type_ of the `\"species\"` column is string. Ideally, we want it to be categorical, so that there's no confusion about the fact that we're dealing with a finite number of mutually exclusive groups here. Another advantage is that it allows plotting tools to realise what kind of data it is working with.\n",
"\n",
- "We can transform the variable to a categorical variable using **pandas** like so:"
+ "We can transform the variable to a categorical variable using **polars** like so:\n"
]
},
{
@@ -614,7 +615,7 @@
"metadata": {},
"outputs": [],
"source": [
- "penguins[\"species\"] = penguins[\"species\"].astype(\"category\")\n",
+ "penguins = penguins.cast({\"species\": pl.Categorical})\n",
"penguins.head()"
]
},
@@ -623,7 +624,7 @@
"id": "06d834a5",
"metadata": {},
"source": [
- "You will learn more about categorical variables later in the book."
+ "You will learn more about categorical variables later in the book.\n"
]
},
{
@@ -631,12 +632,11 @@
"id": "f9ca3124",
"metadata": {},
"source": [
- "\n",
"### A numerical variable\n",
"\n",
"A variable is **numerical** (or quantitative) if it can take on a wide range of numerical values, and it is sensible to add, subtract, or take averages with those values. Numerical variables can be continuous or discrete.\n",
"\n",
- "One commonly used visualisation for distributions of continuous variables is a histogram."
+ "One commonly used visualisation for distributions of continuous variables is a histogram.\n"
]
},
{
@@ -661,7 +661,7 @@
"You should always explore a variety of binwidths when working with histograms, as different binwidths can reveal different patterns.\n",
"In the plots below a binwidth of 20 is too narrow, resulting in too many bars, making it difficult to determine the shape of the distribution.\n",
"Similarly, a binwidth of 2,000 is too high, resulting in all data being binned into only three bars, and also making it difficult to determine the shape of the distribution.\n",
- "A binwidth of 200 provides a sensible balance, but you should always look at your data a few different ways, especially with histograms as they can be misleading."
+ "A binwidth of 200 provides a sensible balance, but you should always look at your data a few different ways, especially with histograms as they can be misleading.\n"
]
},
{
@@ -710,7 +710,7 @@
" geom_bar(fill = \"red\"))\n",
" ```\n",
"\n",
- "3. What does the `bins` argument in `geom_histogram()` do?"
+ "3. What does the `bins` argument in `geom_histogram()` do?\n"
]
},
{
@@ -722,7 +722,7 @@
"\n",
"To visualise a relationship we need to have at least two variables mapped to aesthetics of a plot—though you should remember that correlation is not causation, and causation is not correlation!\n",
"\n",
- "In the following sections you will learn about commonly used plots for visualising relationships between two or more variables and the geoms used for creating them."
+ "In the following sections you will learn about commonly used plots for visualising relationships between two or more variables and the geoms used for creating them.\n"
]
},
{
@@ -738,17 +738,16 @@
"\n",
"It is also useful for identifying potential outliers. Each boxplot consists of:\n",
"\n",
- "- A box that indicates the range of the middle half of the data, a distance known as the interquartile range (IQR), stretching from the 25th percentile of the distribution to the 75th percentile.\n",
- " In the middle of the box is a line that displays the median, i.e. 50th percentile, of the distribution.\n",
- " These three lines give you a sense of the spread of the distribution and whether or not the distribution is symmetric about the median or skewed to one side.\n",
- "\n",
- "- Visual points that display observations that fall more than 1.5 times the IQR from either edge of the box.\n",
- " These outlying points are unusual so are plotted individually.\n",
+ "- A box that indicates the range of the middle half of the data, a distance known as the interquartile range (IQR), stretching from the 25th percentile of the distribution to the 75th percentile.\n",
+ " In the middle of the box is a line that displays the median, i.e. 50th percentile, of the distribution.\n",
+ " These three lines give you a sense of the spread of the distribution and whether or not the distribution is symmetric about the median or skewed to one side.\n",
"\n",
- "- A line (or whisker) that extends from each end of the box and goes to the farthest non-outlier point in the distribution.\n",
+ "- Visual points that display observations that fall more than 1.5 times the IQR from either edge of the box.\n",
+ " These outlying points are unusual so are plotted individually.\n",
"\n",
+ "- A line (or whisker) that extends from each end of the box and goes to the farthest non-outlier point in the distribution.\n",
"\n",
- "Let's take a look at the distribution of body mass by species using `geom_boxplot()`:"
+ "Let's take a look at the distribution of body mass by species using `geom_boxplot()`:\n"
]
},
{
@@ -766,7 +765,7 @@
"id": "97b24caa",
"metadata": {},
"source": [
- "Alternatively, we can make probability density plots with `geom_density()`."
+ "Alternatively, we can make probability density plots with `geom_density()`.\n"
]
},
{
@@ -788,7 +787,7 @@
"\n",
"Additionally, we can map `species` to both `color` and `fill` aesthetics and use the `alpha` aesthetic to add transparency to the filled density curves.\n",
"This aesthetic takes values between 0 (completely transparent) and 1 (completely opaque).\n",
- "In the following plot it's *set* to 0.5."
+ "In the following plot it's _set_ to 0.5.\n"
]
},
{
@@ -811,8 +810,8 @@
"source": [
"Note the terminology we have used here:\n",
"\n",
- "- We *map* variables to aesthetics if we want the visual attribute represented by that aesthetic to vary based on the values of that variable.\n",
- "- Otherwise, we *set* the value of an aesthetic.\n"
+ "- We _map_ variables to aesthetics if we want the visual attribute represented by that aesthetic to vary based on the values of that variable.\n",
+ "- Otherwise, we _set_ the value of an aesthetic.\n"
]
},
{
@@ -829,7 +828,7 @@
"The first plot shows the frequencies of each species of penguins on each island.\n",
"The plot of frequencies show that there are equal numbers of Adelies on each island.\n",
"\n",
- "But we don't have a good sense of the percentage balance within each island."
+ "But we don't have a good sense of the percentage balance within each island.\n"
]
},
{
@@ -867,7 +866,7 @@
"id": "cc83c3db",
"metadata": {},
"source": [
- "In creating these bar charts, we map the variable that will be separated into bars to the `x` aesthetic, and the variable that will change the colors inside the bars to the `fill` aesthetic."
+ "In creating these bar charts, we map the variable that will be separated into bars to the `x` aesthetic, and the variable that will change the colors inside the bars to the `fill` aesthetic.\n"
]
},
{
@@ -927,7 +926,7 @@
"\n",
"To facet your plot by a single variable, use `facet_wrap()`.\n",
"\n",
- "The first argument of `facet_wrap()` tells the function what variable to have in successive charts. The variable that you pass to `facet_wrap()` should be categorical."
+ "The first argument of `facet_wrap()` tells the function what variable to have in successive charts. The variable that you pass to `facet_wrap()` should be categorical.\n"
]
},
{
@@ -949,7 +948,7 @@
"id": "ee5a3eed",
"metadata": {},
"source": [
- "You will learn about many other geoms for visualising distributions of variables and relationships between them in later chapters."
+ "You will learn about many other geoms for visualising distributions of variables and relationships between them in later chapters.\n"
]
},
{
@@ -971,7 +970,7 @@
" ggplot(\n",
" data = penguins,\n",
" mapping = aes(\n",
- " x = \"bill_length_mm\", y = \"bill_depth_mm\", \n",
+ " x = \"bill_length_mm\", y = \"bill_depth_mm\",\n",
" color = \"species\", shape = \"species\"\n",
" )\n",
" ) +\n",
@@ -1023,9 +1022,9 @@
"source": [
"This saved the figure to disk at the location shown—by default it's in a subdirectory called \"lets-plot-images\".\n",
"\n",
- "We used the file format \"svg\". There are lots of output options to choose from to save your file to. Remember that, for graphics, *vector formats* are generally better than *raster formats*. In practice, this means saving plots in svg or pdf formats over jpg or png file formats. The svg format works in a lot of contexts (including Microsoft Word) and is a good default. To choose between formats, just supply the file extension and the file type will change automatically, eg \"chart.svg\" for svg or \"chart.png\" for png. You can also save figures in HTML format.\n",
+ "We used the file format \"svg\". There are lots of output options to choose from to save your file to. Remember that, for graphics, _vector formats_ are generally better than _raster formats_. In practice, this means saving plots in svg or pdf formats over jpg or png file formats. The svg format works in a lot of contexts (including Microsoft Word) and is a good default. To choose between formats, just supply the file extension and the file type will change automatically, eg \"chart.svg\" for svg or \"chart.png\" for png. You can also save figures in HTML format.\n",
"\n",
- "If you're using a raster format then you'll need to specify how big the figure is via the *scale* keyword argument."
+ "If you're using a raster format then you'll need to specify how big the figure is via the _scale_ keyword argument.\n"
]
},
{
@@ -1051,7 +1050,7 @@
"source": [
"### Exercises\n",
"\n",
- "1. Save the figure above as a PNG. Try varying the scale."
+ "1. Save the figure above as a PNG. Try varying the scale.\n"
]
},
{
@@ -1066,13 +1065,12 @@
"We have all been writing Python code for years, but every day we still write code that doesn't work on the first try!\n",
"\n",
"Start by carefully comparing the code that you're running to the code in the book: A misplaced character can make all the difference!\n",
- "Make sure that every `(` is matched with a `)` and every `\"` is paired with another `\"`. In Visual Studio Code, you can get extensions that colour match brackets so you can easily see if you closed them or not.\n",
+ "Make sure that every `(` is matched with a `)` and every `\"` is paired with another `\"`. In Visual Studio Code, you can get extensions that colour match brackets so you can easily see if you closed them or not.\n",
"\n",
"Sometimes you'll run the code and nothing happens.\n",
"\n",
"For those coming from the R statistical programming language, you may be concerned about getting your `+` in the wrong place. Have no fear, however, as in the syntax for **letsplot** the `+` can go at the start or the end of the line.\n",
"\n",
- "\n",
"If you're still stuck, try the help.\n",
"You can get help about any Python function by running `help(function_name)` in the interactive window.\n",
"Don't worry if the help doesn't seem that helpful - instead skip down to the examples and look for code that matches what you're trying to do.\n",
@@ -1095,7 +1093,7 @@
"We'll use visualisations again and again throughout this book, introducing new techniques as we need them as well as do a deeper dive into creating visualisations with **letsplot** in subsequent chapters.\n",
"\n",
"With the basics of visualisation under your belt, in the next chapter we're going to switch gears a little and give you some practical workflow advice.\n",
- "We intersperse workflow advice with data science tools throughout this part of the book because it'll help you stay organised as you write more Python code."
+ "We intersperse workflow advice with data science tools throughout this part of the book because it'll help you stay organised as you write more Python code.\n"
]
}
],
@@ -1107,7 +1105,7 @@
"main_language": "python"
},
"kernelspec": {
- "display_name": ".venv",
+ "display_name": "python4ds",
"language": "python",
"name": "python3"
},
diff --git a/data/bake_sale.xlsx b/data/bake_sale.xlsx
index cf475a2..e122900 100644
Binary files a/data/bake_sale.xlsx and b/data/bake_sale.xlsx differ
diff --git a/databases.ipynb b/databases.ipynb
index f14914c..661cfd1 100644
--- a/databases.ipynb
+++ b/databases.ipynb
@@ -17,7 +17,7 @@
"\n",
"### Prerequisites\n",
"\n",
- "You will need the **pandas**, **SQLModel**, and **ibis** packages for this chapter. You probably already have **pandas** installed; to install **SQLModel** and **ibis** respectively run `uv add sqlmodel` and `uv add ibis-framework` on your computer's command line. First, let's bring in some general packages and turn off verbose warnings."
+ "You will need the **polars**, **SQLModel**, and **ibis** packages for this chapter. You probably already have **polars** installed; to install **SQLModel** and **ibis** respectively run `uv add sqlmodel` and `uv add ibis-framework` on your computer's command line. First, let's bring in some general packages and turn off verbose warnings."
]
},
{
@@ -39,10 +39,9 @@
"metadata": {},
"source": [
"## Database Basics\n",
- "\n",
- "At the simplest level, you can think about a database as a collection of data frames, called **tables** in database terminology.\n",
- "Like a **pandas** data frame, a database table is a collection of named columns, where every value in the column is the same type.\n",
- "There are three high level differences between data frames and database tables:\n",
+ "At the simplest level, you can think about a database as a collection of data frames, called **tables** in database terminology. \n",
+ "Like a **Polars** DataFrame, a database table is a collection of named columns, where every value in a column shares the same data type. \n",
+ "There are three high-level differences between data frames and database tables:\n",
"\n",
"- Database tables are stored on disk (ie on file) and can be arbitrarily large.\n",
" Data frames are stored in memory, and are fundamentally limited (although that limit is still big enough for many problems). You can think about the difference between on disk and in memory as being like the difference between long-term and short-term memory (and you have much more limited capacity in the latter).\n",
@@ -68,7 +67,7 @@
"\n",
"- You'll always use a database interface that provides a connection to the database, for example Python's built-in **sqlite** package\n",
"\n",
- "- You'll also use a package that pushes and/or pulls data to/from the database, for example **pandas**\n",
+ "- You'll also use a package that pushes and/or pulls data to/from the database, for example **polars**\n",
"\n",
"The precise details of the connection varies a lot from DBMS to DBMS so unfortunately we can't cover all the details here. The initial setup will often take a little fiddling (and maybe some research) to get right, but you'll generally only need to do it once. We'll do the best we can to cover some basics here.\n",
"\n",
@@ -112,7 +111,7 @@
"id": "2992b718",
"metadata": {},
"source": [
- "Note that the output here is in the form a Python object called a tuple. If we wanted to put this into a **pandas** data frame, we can just pass it straight in:"
+ "Note that the output here is in the form of a Python object called a tuple. If we want to convert this into a **Polars** DataFrame, we can pass it to `pl.DataFrame()`. When working with tuples, you may need to provide column names using the **schema** argument or specify **orient=\"row\"** so Polars correctly interprets the structure."
]
},
{
@@ -122,9 +121,11 @@
"metadata": {},
"outputs": [],
"source": [
- "import pandas as pd\n",
+ "import polars as pl\n",
+ "\n",
+ "df = pl.DataFrame(rows, orient=\"row\")\n",
"\n",
- "pd.DataFrame(rows)"
+ "df"
]
},
{
@@ -316,9 +317,9 @@
"source": [
"### Joins\n",
"\n",
- "If you're familiar with joins in **pandas**, SQL joins are very similar. Let's see if we can join the 'album' and 'track' tables to find the *name* of the albums in the above query.\n",
+ "If you’re familiar with joins in **polars**, SQL joins are very similar. Let’s see if we can join the 'album' and 'track' tables to find the *name* of the albums in the above query.\n",
"\n",
- "Note that as soon as we have the *same* column names in more than one table, we need to specify the table we are referring to when we use that column name. There are different options for joins (eg `INNER`, `LEFT`) that you can find out more about [here](https://en.wikipedia.org/wiki/Join_(SQL)).\n"
+ "In polars, you use the `df.join()` method, which defaults to an \"inner\" join. Note that if you have the same column names in both tables, Polars will often append a suffix (like _right) to the duplicate names to keep them distinct, unless you specify otherwise. There are different options for joins (eg `INNER`, `LEFT`) that you can find out more about [here](https://en.wikipedia.org/wiki/Join_(SQL)).\n"
]
},
{
@@ -403,9 +404,9 @@
"id": "495f97e5",
"metadata": {},
"source": [
- "## SQL with **pandas**\n",
+ "## SQL with **polars**\n",
"\n",
- "**pandas** is well-equipped for working with SQL. We can simply push the query we just created straight through using its `read_sql()` function—but bear in mind we need to pass in the connection we created to the database too:"
+ "**polars** is well-equipped for working with SQL. We can simply push the query we just created straight through using its `read_database()` function—but bear in mind we need to pass in the connection we created to the database too:"
]
},
{
@@ -415,7 +416,10 @@
"metadata": {},
"outputs": [],
"source": [
- "pd.read_sql(sql_join, con)"
+ "df = pl.read_database(\n",
+ " query=sql_join, # your SQL query (string)\n",
+ " connection=con, # your connection object (SQLAlchemy, psycopg2 cursor, etc.)\n",
+ ")"
]
},
{
@@ -435,7 +439,7 @@
"source": [
"## SQL with **ibis**\n",
"\n",
- "It's not exactly satisfactory to have to write out your SQL queries in text. What if we could create commands directly from **pandas** commands? You can't *quite* do that, but there's a package that gets you pretty close and it's called [**ibis**](https://ibis-project.org/). **ibis** is particularly useful when you are reading from a database and want to query it just like you would a **pandas** data frame.\n",
+ "It's not exactly satisfactory to have to write out your SQL queries in text. What if we could create commands directly from **polars** commands? You can't *quite* do that, but there's a package that gets you pretty close and it's called [**ibis**](https://ibis-project.org/). **ibis** is particularly useful when you are reading from a database and want to query it just like you would a **polars** data frame.\n",
"\n",
"**Ibis** can connect to local databases (eg a SQLite database), server-based databases (eg Postgres), or cloud-based databased (eg Google's BigQuery). The syntax to make a connection is, for example, `ibis.bigquery.connect`.\n",
"\n",
@@ -462,7 +466,7 @@
"id": "6dcd7d71",
"metadata": {},
"source": [
- "Okay, now let's reproduce the following query: \"SELECT albumid, AVG(milliseconds)/1e3/60 FROM track GROUP BY albumid ORDER BY AVG(milliseconds) ASC LIMIT 5;\". We'll use a groupby, a mutate (which you can think of like **pandas**' assign statement), a sort, and then `limit()` to only show the first five entries."
+ "Okay, now let's reproduce the following query: \"SELECT albumid, AVG(milliseconds)/1e3/60 FROM track GROUP BY albumid ORDER BY AVG(milliseconds) ASC LIMIT 5;\". We'll use a group_by, a mutate (which you can think of like **polars** assign statement), a sort, and then `limit()` to only show the first five entries."
]
},
{
diff --git a/dataframe_illustration.svg b/dataframe_illustration.svg
new file mode 100644
index 0000000..2c35021
--- /dev/null
+++ b/dataframe_illustration.svg
@@ -0,0 +1,233 @@
+
+
diff --git a/index.md b/index.md
index 965c550..6086806 100644
--- a/index.md
+++ b/index.md
@@ -5,29 +5,14 @@ aliases:
# Welcome
-[](https://zenodo.org/doi/10.5281/zenodo.10518241) 
+This is the website for **Python for Data Science**, a book heavily inspired by the excellent [**R for Data Science (2e)**](https://r4ds.hadley.nz/). This book will teach you how to load up, transform, visualize, and begin to understand your data. The book aims to give you the skills you need to code for data science. It's suitable for people who have some familiarity with the ideas behind programming and coding but who don't yet know how to do data science.
-This is the website for **Python for Data Science**, a book heavily inspired by the excellent [**R for Data Science (2e)**](https://r4ds.hadley.nz/). This book will teach you how to load up, transform, visualise, and begin to understand your data. The book aims to give you the skills you need to code for data science. It's suitable for people who have some familiarity with the ideas behind programming and coding but who don't yet know how to do data science.
+This book teaches you how to do data science using one of the world's most popular programming languages, Python. While Python is a general-purpose language, meaning it is used for a wide range of tasks, it is also the most widely used language for data science (though note that both SQL and R are also used for data science).
-This book teaches you how to do data science using one of the world's most popular programming languages, Python. While Python is a general purpose language, which means it is used for a wide range of tasks, it is also the most widely used language for data science (although note that both SQL and R are also used for data science).
+__This [fork](https://github.com/aeturrell/python4DS) of a [fork](https://github.com/hadley/r4ds) focuses on Polars instead of Pandas for data wrangling.__
To begin your data science journey, head to the next page.
## Contributors to Python4DS
Contributing is very much encouraged. If you're looking for content to implement or tweak, we aim to follow the structure and content of **R for Data Science (2e)** and you can find open [issues here](https://github.com/aeturrell/python4DS/issues). For larger contributions of content, it's probably best to check with other contributors first.
-
-We thank the following contributors:
-
-- [Arthur Turrell](https://aeturrell.com/), who has also contributed to [*Coding for Economists*](https://aeturrell.github.io/coding-for-economists) and wrote popular non-fiction book [*The Star Builders*](https://aeturrell.com/thestarbuilders/thestarbuilders.html)
-- [Pietro Monticone](https://github.com/pitmonticone)
-- [Antonio Mele](https://github.com/meleantonio)
-- [Igor Alshannikov](https://github.com/alshan)
-- [Umair Durrani](https://github.com/durraniu)
-- [Zeki Akyol](https://github.com/zekiakyol)
-- [Yiben Huang](https://github.com/yibenhuang)
-- [William Chiu](https://github.com/crossxwill)
-- [udurraniAtPresage](https://github.com/udurraniAtPresage)
-- [Josh Holman](https://github.com/TheJolman)
-- [Kenytt Avery](https://github.com/ProfAvery)
-- [Bradley Phipps](https://github.com/hotshotberad)
diff --git a/pyproject.toml b/pyproject.toml
index 016e885..7740186 100644
--- a/pyproject.toml
+++ b/pyproject.toml
@@ -1,11 +1,12 @@
[project]
name = "python4ds"
-version = "1.0.5"
+version = "0.0.1"
description = "The online book that teaches you how to use Python for data science."
readme = "README.md"
requires-python = ">=3.12.0,<3.13"
dependencies = [
"beautifulsoup4>=4.12.3",
+ "fastexcel>=0.19.0",
"graphviz>=0.20.3",
"ibis-framework[sqlite]>=9.5.0",
"ipykernel>=6.29.5",
@@ -36,6 +37,7 @@ dependencies = [
"toml>=0.10.2",
"watermark>=2.5.0",
"wbgapi>=1.0.14",
+ "xlsxwriter>=3.2.0",
"yfinance>=1.2.1",
]
diff --git a/rectangling.ipynb b/rectangling.ipynb
index f0bcf47..e565c1d 100644
--- a/rectangling.ipynb
+++ b/rectangling.ipynb
@@ -41,7 +41,7 @@
"source": [
"### Prerequisites\n",
"\n",
- "This chapter will use the **pandas** data analysis package."
+ "This chapter will use the **polars** data analysis package.\n"
]
},
{
@@ -51,7 +51,7 @@
"source": [
"## Lists\n",
"\n",
- "Lists are a really useful way to work with lots of data at once. They're defined with square brackets, with entries separated by commas. "
+ "Lists are a really useful way to work with lots of data at once. They're defined with square brackets, with entries separated by commas.\n"
]
},
{
@@ -70,7 +70,7 @@
"id": "29b10d07",
"metadata": {},
"source": [
- "You can also construct them by appending entries:"
+ "You can also construct them by appending entries:\n"
]
},
{
@@ -89,7 +89,7 @@
"id": "d8d4f6ed",
"metadata": {},
"source": [
- "And you can access earlier entries using an index, which begins at 0 and ends at one less than the length of the list (this is the convention in many programming languages). For instance, to print specific entries at the start, using `0`, and end, using `-1`:"
+ "And you can access earlier entries using an index, which begins at 0 and ends at one less than the length of the list (this is the convention in many programming languages). For instance, to print specific entries at the start, using `0`, and end, using `-1`:\n"
]
},
{
@@ -110,7 +110,7 @@
"source": [
"::: {.callout-tip title=\"Exercise\"}\n",
"How might you access the penultimate entry in a list object if you didn't know how many elements it had?\n",
- ":::"
+ ":::\n"
]
},
{
@@ -118,7 +118,7 @@
"id": "6aea9157",
"metadata": {},
"source": [
- "As well as accessing positions in lists using indexing, you can use *slices* on lists. This uses the colon character, `:`, to stand in for 'from the beginning' or 'until the end' (when only appearing once). For instance, to print just the last two entries, we would use the index `-2:` to mean from the second-to-last onwards. Here are two distinct examples: getting the first three and last three entries to be successively printed:"
+ "As well as accessing positions in lists using indexing, you can use _slices_ on lists. This uses the colon character, `:`, to stand in for 'from the beginning' or 'until the end' (when only appearing once). For instance, to print just the last two entries, we would use the index `-2:` to mean from the second-to-last onwards. Here are two distinct examples: getting the first three and last three entries to be successively printed:\n"
]
},
{
@@ -137,7 +137,7 @@
"id": "c82b5c4a",
"metadata": {},
"source": [
- "Slicing can be even more elaborate than that because we can jump entries using a second colon. Here's a full example that begins at the second entry (remember the index starts at 0), runs up until the second-to-last entry (exclusive), and jumps every other entry inbetween (range just produces a list of integers from the value to one less than the last):"
+ "Slicing can be even more elaborate than that because we can jump entries using a second colon. Here's a full example that begins at the second entry (remember the index starts at 0), runs up until the second-to-last entry (exclusive), and jumps every other entry inbetween (range just produces a list of integers from the value to one less than the last):\n"
]
},
{
@@ -159,7 +159,7 @@
"id": "813e09bc",
"metadata": {},
"source": [
- "A handy trick is that you can print a reversed list entirely using double colons:"
+ "A handy trick is that you can print a reversed list entirely using double colons:\n"
]
},
{
@@ -179,7 +179,7 @@
"source": [
"::: {.callout-tip title=\"Exercise\"}\n",
"Slice the `list_example` from earlier to get only the first five entries.\n",
- ":::"
+ ":::\n"
]
},
{
@@ -187,7 +187,7 @@
"id": "b6ff3ca4",
"metadata": {},
"source": [
- "What's amazing about lists is that they can hold any type, including other lists! Here's a valid example of a list that's got a lot going on:"
+ "What's amazing about lists is that they can hold any type, including other lists! Here's a valid example of a list that's got a lot going on:\n"
]
},
{
@@ -217,7 +217,7 @@
"source": [
"### Hierarchical Data in Lists\n",
"\n",
- "Because lists can contain more lists (and so on), they can be used to put hierachical data in. Let's take a look at an example:"
+ "Because lists can contain more lists (and so on), they can be used to put hierachical data in. Let's take a look at an example:\n"
]
},
{
@@ -236,7 +236,7 @@
"id": "57a81b53",
"metadata": {},
"source": [
- "Now, say we wanted to reduce this to a single list. We can do it with a *list comprehension*:"
+ "Now, say we wanted to reduce this to a single list. We can do it with a _list comprehension_:\n"
]
},
{
@@ -254,7 +254,7 @@
"id": "8e96185a",
"metadata": {},
"source": [
- "What we're saying here is take all of the values of every little list and put them into a single list."
+ "What we're saying here is take all of the values of every little list and put them into a single list.\n"
]
},
{
@@ -264,7 +264,7 @@
"source": [
"### From Lists to Data Frames\n",
"\n",
- "Occassionally, you'll have data in lists that you wish to turn into a data frame. For example, perhaps you have a list of lists like this:"
+ "Occassionally, you'll have data in lists that you wish to turn into a data frame. For example, perhaps you have a list of lists like this:\n"
]
},
{
@@ -282,7 +282,7 @@
"id": "fcfc2d3c",
"metadata": {},
"source": [
- "You can pass this straight into a constructor for a data frame as the `data=` keyword argument (adding in other info as necessary). Note that this is four lists of three entries, so the inner loop has entries in 0 to 2... it is this inner loop that will be used as the *rows* of any data frame with the number of entries in each inner list equal to the number of *columns*."
+ "You can pass this straight into a constructor for a data frame as the `data=` keyword argument (adding in other info as necessary). Note that this is four lists of three entries, so the inner loop has entries in 0 to 2... it is this inner loop that will be used as the _rows_ of any data frame with the number of entries in each inner list equal to the number of _columns_.\n"
]
},
{
@@ -292,9 +292,9 @@
"metadata": {},
"outputs": [],
"source": [
- "import pandas as pd\n",
+ "import polars as pl\n",
"\n",
- "pd.DataFrame(data=list_of_lists, columns=[\"a\", \"b\", \"c\"])"
+ "df = pl.DataFrame(data=list_of_lists, schema=[\"a\", \"b\", \"c\", \"d\"])"
]
},
{
@@ -302,7 +302,7 @@
"id": "cc797c89",
"metadata": {},
"source": [
- "There's one more trick to show you: explode. This is useful when you have data that has more than one level of list depth. Let's say you read in some data with a complex hierarchical structure like this:"
+ "There's one more trick to show you: explode. This is useful when you have data that has more than one level of list depth. Let's say you read in some data with a complex hierarchical structure like this:\n"
]
},
{
@@ -312,12 +312,13 @@
"metadata": {},
"outputs": [],
"source": [
- "df = pd.DataFrame(\n",
+ "df = pl.DataFrame(\n",
" {\n",
- " \"alpha\": [[0, 1, 2], \"foo\", [], [3, 4]],\n",
- " \"beta\": 1,\n",
- " \"gamma\": [[\"a\", \"b\", \"c\"], pd.NA, [], [\"d\", \"e\"]],\n",
- " }\n",
+ " \"alpha\": [[\"0,1,2\"], \"foo\", [], [\"3,4\"]],\n",
+ " \"beta\": [1, 1, 1, 1],\n",
+ " \"gamma\": [[\"a\", \"b\", \"c\"], None, [], [\"d\", \"e\"]],\n",
+ " },\n",
+ " strict=False,\n",
")\n",
"df"
]
@@ -327,7 +328,7 @@
"id": "91bb97aa",
"metadata": {},
"source": [
- "We have multiple rows and columns that contain lists. In some situations, it's fine to have a list in a column but here it's probably not as it's mixed in with other types of data. We can use `explode()` to split out the columns further length-wise"
+ "We have multiple rows and columns that contain lists. In some situations, it's fine to have a list in a column but here it's probably not as it's mixed in with other types of data. We can use `explode()` to split out the columns further length-wise\n"
]
},
{
@@ -337,7 +338,7 @@
"metadata": {},
"outputs": [],
"source": [
- "df.explode(\"alpha\")"
+ "df.explode(\"gamma\")"
]
},
{
@@ -352,7 +353,7 @@
"The table below compares the different data types found in Python and JSON.\n",
"\n",
"| JSON OBJECT | PYTHON OBJECT |\n",
- "|---------------|---------------|\n",
+ "| ------------- | ------------- |\n",
"| object | dict |\n",
"| array | list |\n",
"| string | str |\n",
@@ -362,9 +363,9 @@
"| true | True |\n",
"| false | False |\n",
"\n",
- "There are typically two operations you may want to do with JSON data: 1) turn JSON data in a Python object (eg JSON to Python dictionary) or vice versa (known as deserialisation and serialisation respectively); and 2) converting a deserialised object into a *different* kind of Python object.\n",
+ "There are typically two operations you may want to do with JSON data: 1) turn JSON data in a Python object (eg JSON to Python dictionary) or vice versa (known as deserialisation and serialisation respectively); and 2) converting a deserialised object into a _different_ kind of Python object.\n",
"\n",
- "Let's look at each in turn."
+ "Let's look at each in turn.\n"
]
},
{
@@ -378,7 +379,7 @@
"\n",
"#### From the Web\n",
"\n",
- "We'll get some JSON data from an API. Let's grab the latest UK unemployment data (timeseries code \"MGSX\" and dataset code \"LMS\")."
+ "We'll get some JSON data from an API. Let's grab the latest UK unemployment data (timeseries code \"MGSX\" and dataset code \"LMS\").\n"
]
},
{
@@ -401,7 +402,7 @@
"id": "051d3b4a",
"metadata": {},
"source": [
- "Let's check what type we got:"
+ "Let's check what type we got:\n"
]
},
{
@@ -421,7 +422,7 @@
"source": [
"As expected, the JSON data has automatically been read in as a dictionary—but be wary that the fields have been read in as text rather than numbers, datetimes, and other specific data types.\n",
"\n",
- "We could print the whole object out but that would take up a lot of space; instead let's look at a couple of entries under the \"months\" key."
+ "We could print the whole object out but that would take up a lot of space; instead let's look at a couple of entries under the \"months\" key.\n"
]
},
{
@@ -441,7 +442,7 @@
"source": [
"#### From a File or Stream\n",
"\n",
- "For this exercise, you'll need to download the JSON file 'cakes.json' from the [data folder of the repository](https://github.com/aeturrell/python4DS/tree/main/data) associated with this book and save it in a sub-folder called \"data\". We can take a peek at the data using the terminal (which is what the preceeding exclamation mark means):"
+ "For this exercise, you'll need to download the JSON file 'cakes.json' from the [data folder of the repository](https://github.com/aeturrell/python4DS/tree/main/data) associated with this book and save it in a sub-folder called \"data\". We can take a peek at the data using the terminal (which is what the preceeding exclamation mark means):\n"
]
},
{
@@ -467,7 +468,7 @@
"id": "0c664ab6",
"metadata": {},
"source": [
- "We use the built-in **json** library to read this into Python (you could also use a file path here—more on how in a moment):"
+ "We use the built-in **json** library to read this into Python (you could also use a file path here—more on how in a moment):\n"
]
},
{
@@ -488,7 +489,7 @@
"id": "df41f92b",
"metadata": {},
"source": [
- "Note that not everything is the same in going from JSON text to a Python dictionary: JSON uses `null` rather than `None`, won't accept trailing commas at the end of lists, and has basic types that are lists, strings (and all keys must be strings), numbers, booleans, and nulls. Let's now see how to write a Python dictionary back to a JSON, perhaps for writing to file:"
+ "Note that not everything is the same in going from JSON text to a Python dictionary: JSON uses `null` rather than `None`, won't accept trailing commas at the end of lists, and has basic types that are lists, strings (and all keys must be strings), numbers, booleans, and nulls. Let's now see how to write a Python dictionary back to a JSON, perhaps for writing to file:\n"
]
},
{
@@ -507,7 +508,7 @@
"id": "5f9445b8",
"metadata": {},
"source": [
- "To write to a file, you would use the pattern:"
+ "To write to a file, you would use the pattern:\n"
]
},
{
@@ -518,7 +519,7 @@
"```python\n",
"with open('data/json_data_output.json', 'w') as outfile:\n",
" json.dump(json_stream, outfile)\n",
- "```"
+ "```\n"
]
},
{
@@ -530,7 +531,7 @@
"\n",
"```python\n",
"json.load(open(\"data/json_data_output.json\"))\n",
- "```"
+ "```\n"
]
},
{
@@ -540,7 +541,7 @@
"source": [
"### From JSON data to Data Frame\n",
"\n",
- "**pandas** has lots of options for turning JSON or dictionary data into a data frame. You do need to think a little bit about the structure of the data underneath though:\n"
+ "**polars** has lots of options for turning JSON or dictionary data into a data frame. You do need to think a little bit about the structure of the data underneath though:\n"
]
},
{
@@ -550,9 +551,9 @@
"metadata": {},
"outputs": [],
"source": [
- "import pandas as pd\n",
+ "import polars as pl\n",
"\n",
- "pd.DataFrame(result[\"toppings\"], columns=[\"id\", \"type\"])"
+ "df = pl.DataFrame(result[\"toppings\"], schema=[\"id\", \"type\"])"
]
},
{
@@ -560,7 +561,7 @@
"id": "a1346020",
"metadata": {},
"source": [
- "The web-scraped data we downloaded earlier had a more complicated structure, but **pandas** has a `json_normalize()` function that can cope with this. For example, with the following data, there are many missing entries but `json_normalize()` can still parse it into a Data Frame."
+ "The web-scraped data we downloaded earlier had a more complicated structure, but **polars** has a `json_normalize()` function that can cope with this. For example, with the following data, there are many missing entries but `json_normalize()` can still parse it into a Data Frame.\n"
]
},
{
@@ -575,7 +576,7 @@
" {\"name\": {\"given\": \"Mark\", \"family\": \"Regner\"}},\n",
" {\"id\": 2, \"name\": \"Faye Raker\"},\n",
"]\n",
- "pd.json_normalize(data)"
+ "pl.json_normalize(data)"
]
},
{
@@ -583,7 +584,7 @@
"id": "7eaf00e1",
"metadata": {},
"source": [
- "And we can control the level that properties like 'name' are split out to as well (you can check out more options over at the [**pandas** documentation](https://pandas.pydata.org/docs/reference/api/pandas.json_normalize.html))"
+ "And we can control the level that properties like 'name' are split out to as well (you can check out more options over at the [**polars** documentation](https://docs.pola.rs/api/python/stable/reference/api/polars.json_normalize.html))\n"
]
},
{
@@ -593,7 +594,7 @@
"metadata": {},
"outputs": [],
"source": [
- "pd.json_normalize(data, max_level=0)"
+ "pl.json_normalize(data, max_level=0)"
]
},
{
@@ -601,7 +602,7 @@
"id": "78d637e5",
"metadata": {},
"source": [
- "As well as the JSON normalise function, **pandas** has a `from_dict()` method to work with simpler dictionary objects."
+ "As well as the JSON normalise function, **polars** has a `from_dict()` method to work with simpler dictionary objects.\n"
]
}
],
@@ -613,7 +614,7 @@
"main_language": "python"
},
"kernelspec": {
- "display_name": "Python 3 (ipykernel)",
+ "display_name": "python4ds",
"language": "python",
"name": "python3"
},
diff --git a/spreadsheets.ipynb b/spreadsheets.ipynb
index 12f1672..1c1f378 100644
--- a/spreadsheets.ipynb
+++ b/spreadsheets.ipynb
@@ -11,7 +11,7 @@
"\n",
"This chapter will show you how to work with spreadsheets, for example Microsoft Excel files, in Python. We already saw how to import csv (and tsv) files in @sec-data-import. In this chapter we will introduce you to tools for working with data in Excel spreadsheets and Google Sheets.\n",
"\n",
- "If you or your collaborators are using spreadsheets for organising data that will be ingested by an analytical tool like Python, we recommend reading the paper \"Data Organization in Spreadsheets\" by Karl Broman and Kara Woo {cite}`broman2018data`. The best practices presented in this paper will save you much headache down the line when you import the data from a spreadsheet into Python to analyse and visualise. (For spreadsheets that are meant to be read by humans, we recommend the [good practice tables](https://github.com/best-practice-and-impact/gptables) package.)"
+ "If you or your collaborators are using spreadsheets for organising data that will be ingested by an analytical tool like Python, we recommend reading the paper \"Data Organization in Spreadsheets\" by Karl Broman and Kara Woo {cite}`broman2018data`. The best practices presented in this paper will save you much headache down the line when you import the data from a spreadsheet into Python to analyse and visualise. (For spreadsheets that are meant to be read by humans, we recommend the [good practice tables](https://github.com/best-practice-and-impact/gptables) package.)\n"
]
},
{
@@ -41,7 +41,7 @@
"source": [
"### Prerequisites\n",
"\n",
- "You will need to install the **pandas** package for this chapter. You will also need to install the **openpyxl** package by running `uv add openpyxl` in the terminal."
+ "You will need the **polars** package for this chapter. Install **fastexcel** so `read_excel()` can use the default (fast) engine, **xlsxwriter** for `write_excel()`, and **openpyxl** if you want to use the **openpyxl** engine explicitly (`uv add fastexcel xlsxwriter openpyxl`).\n"
]
},
{
@@ -51,11 +51,11 @@
"source": [
"## Reading Excel (and Similar) Files\n",
"\n",
- "**pandas** can read in xls, xlsx, xlsm, xlsb, odf, ods, and odt files from your local filesystem or from a URL. It also supports an option to read a single sheet or a list of sheets.\n",
+ "**polars** can read in xls, xlsx, xlsm, xlsb, odf, ods, and odt files from your local filesystem or from a URL. It also supports an option to read a single sheet or a list of sheets. The default reader uses **fastexcel** (Rust-backed, via the **fastexcel** Python package); you can also select other engines such as **openpyxl** when you need engine-specific options.\n",
"\n",
"To show how this works, we'll work with an example spreadsheet called \"students.xlsx\". The figure below shows what the spreadsheet looks like.\n",
"\n",
- ""
+ "\n"
]
},
{
@@ -63,7 +63,7 @@
"id": "29f2f4e0",
"metadata": {},
"source": [
- "The first argument to `pd.read_excel()` is the path to the file to read. If you have downloaded the [file]() onto your computer and put it in a subfolder called \"data\" then you would want to use the path \"data/students.xlsx\" but we can also load it directly from the URL."
+ "The first argument to `pl.read_excel()` is the path to the file to read. If you have downloaded the [file]() onto your computer and put it in a subfolder called \"data\" then you would want to use the path \"data/students.xlsx\" but we can also load it directly from the URL.\n"
]
},
{
@@ -73,10 +73,10 @@
"metadata": {},
"outputs": [],
"source": [
- "import pandas as pd\n",
+ "import polars as pl\n",
"\n",
- "students = pd.read_excel(\n",
- " \"https://github.com/aeturrell/python4DS/raw/main/data/students.xlsx\"\n",
+ "students = pl.read_excel(\n",
+ " \"data/students.xlsx\",\n",
")\n",
"students"
]
@@ -88,7 +88,7 @@
"source": [
"We have six students in the data and five variables on each student. However there are a few things we might want to address in this dataset:\n",
"\n",
- "- The column names are all over the place. You can provide column names that follow a consistent format; we recommend `snake_case` using the `names` argument.\n"
+ "- The column names are all over the place. You can rename them to follow a consistent format; we recommend `snake_case`. If you want to replace **every** column name in order, assigning to `students.columns` is clear and short. If you only rename some columns, use `.rename({\"Old Name\": \"new_name\", ...})` with the exact strings from the sheet.\n"
]
},
{
@@ -98,10 +98,14 @@
"metadata": {},
"outputs": [],
"source": [
- "pd.read_excel(\n",
- " \"https://github.com/aeturrell/python4DS/raw/main/data/students.xlsx\",\n",
- " names=[\"student_id\", \"full_name\", \"favourite_food\", \"meal_plan\", \"age\"],\n",
- ")"
+ "students.columns = [\n",
+ " \"student_id\",\n",
+ " \"full_name\",\n",
+ " \"favourite_food\",\n",
+ " \"meal_plan\",\n",
+ " \"age\",\n",
+ "]\n",
+ "students"
]
},
{
@@ -109,8 +113,7 @@
"id": "bb07ad4f",
"metadata": {},
"source": [
- "\n",
- "- `age` is read in as a column of objects, but it really should be numeric. Just like with `read_csv()`, you can supply a `dtype` argument to `read_excel()` and specify the data types for the columns of data you read in. Your options include `\"boolean\"`, `\"int\"`, `\"float\"`, `\"datetime\"`, `\"string\"`, and more. But we can see right away that this isn't going to work with the \"age\" column as it mixes numbers and text: so we first need to map its text to numbers."
+ "- `age` may be inferred as strings (for example **Utf8**) when the column mixes numeric values and text, but we want it numeric. Just like with `read_csv()`, you can supply a `schema_overrides` argument to `read_excel()` and specify Polars data types for the columns you read in (for example `pl.Int64`, `pl.Utf8`, `pl.Boolean`, `pl.Datetime`, and more). That still will not fix a value like `\"five\"` until we map it to a number first.\n"
]
},
{
@@ -120,11 +123,15 @@
"metadata": {},
"outputs": [],
"source": [
- "students = pd.read_excel(\n",
- " \"data/students.xlsx\",\n",
- " names=[\"student_id\", \"full_name\", \"favourite_food\", \"meal_plan\", \"age\"],\n",
- ")\n",
- "students[\"age\"] = students[\"age\"].replace(\"five\", 5)\n",
+ "students = pl.read_excel(\"data/students.xlsx\")\n",
+ "students.columns = [\n",
+ " \"student_id\",\n",
+ " \"full_name\",\n",
+ " \"favourite_food\",\n",
+ " \"meal_plan\",\n",
+ " \"age\",\n",
+ "]\n",
+ "students = students.with_columns(pl.col(\"age\").replace({\"five\": 5}))\n",
"students"
]
},
@@ -133,7 +140,7 @@
"id": "c8a07159",
"metadata": {},
"source": [
- "Okay, now we can apply the data types."
+ "Okay, now we can apply the data types.\n"
]
},
{
@@ -143,16 +150,16 @@
"metadata": {},
"outputs": [],
"source": [
- "students = students.astype(\n",
- " {\n",
- " \"student_id\": \"Int64\",\n",
- " \"full_name\": \"string\",\n",
- " \"favourite_food\": \"string\",\n",
- " \"meal_plan\": \"category\",\n",
- " \"age\": \"Int64\",\n",
- " }\n",
+ "students = students.with_columns(\n",
+ " [\n",
+ " pl.col(\"student_id\").cast(pl.Int64),\n",
+ " pl.col(\"full_name\").cast(pl.Utf8),\n",
+ " pl.col(\"favourite_food\").cast(pl.Utf8),\n",
+ " pl.col(\"meal_plan\").cast(pl.Categorical),\n",
+ " pl.col(\"age\").cast(pl.Int64),\n",
+ " ]\n",
")\n",
- "students.info()"
+ "students.schema"
]
},
{
@@ -160,7 +167,7 @@
"id": "362ff5a5",
"metadata": {},
"source": [
- "It took multiple steps and trial-and-error to load the data in exactly the format we want, and this is not unexpected. Data science is an iterative process. There is no way to know exactly what the data will look like until you load it and take a look at it. The general pattern we used is load the data, take a peek, make adjustments to your code, load it again, and repeat until you're happy with the result."
+ "It took multiple steps and trial-and-error to load the data in exactly the format we want, and this is not unexpected. Data science is an iterative process. There is no way to know exactly what the data will look like until you load it and take a look at it. The general pattern we used is load the data, take a peek, make adjustments to your code, load it again, and repeat until you're happy with the result.\n"
]
},
{
@@ -174,7 +181,7 @@
"\n",
"\n",
"\n",
- "You can read a single sheet using the following command (so as not to show the whole file, we'll use `.head()` to just show the first 5 rows):"
+ "You can read a single sheet using the following command (so as not to show the whole file, we'll use `.head()` to just show the first 5 rows):\n"
]
},
{
@@ -184,8 +191,8 @@
"metadata": {},
"outputs": [],
"source": [
- "pd.read_excel(\n",
- " \"https://github.com/aeturrell/python4DS/raw/main/data/penguins.xlsx\",\n",
+ "pl.read_excel(\n",
+ " \"data/penguins.xlsx\",\n",
" sheet_name=\"Torgersen Island\",\n",
").head()"
]
@@ -195,9 +202,9 @@
"id": "641f6831",
"metadata": {},
"source": [
- "Now this relies on us knowing the names of the sheets in advance. There will be situations where you can to read in data without peeking into the Excel spreadsheet. To read all sheets in, use `sheet_name=None`. The object that's created is a dictionary with key value pairs that are sheet names and data frames respectively. Let's look at the second key value pair (note that we have to convert the keys() and values() objects to list to then retrieve the second element of each using a subscript, ie `list(dictionary.keys())[]`).\n",
+ "Now this relies on us knowing the names of the sheets in advance. There will be situations where you want to read in data without peeking into the Excel spreadsheet. To read all sheets in Polars, use `sheet_id=0` (or `sheet_name=None`, which also works in recent versions of Polars). The object that’s created is a dictionary where the keys are the sheet names and the values are Polars DataFrames. To access a specific sheet, you can convert the keys() or values() to a list and then index into it, ie `list(dictionary.keys())[]` .\n",
"\n",
- "To give a sense of how this works, let's first print all of the retrieved keys:"
+ "To give a sense of how this works, let's first print all of the retrieved keys:\n"
]
},
{
@@ -207,9 +214,9 @@
"metadata": {},
"outputs": [],
"source": [
- "penguins_dict = pd.read_excel(\n",
- " \"https://github.com/aeturrell/python4DS/raw/main/data/penguins.xlsx\",\n",
- " sheet_name=None,\n",
+ "penguins_dict = pl.read_excel(\n",
+ " \"data/penguins.xlsx\",\n",
+ " sheet_id=0,\n",
")\n",
"print([x for x in penguins_dict.keys()])"
]
@@ -219,7 +226,7 @@
"id": "076f1ebe",
"metadata": {},
"source": [
- "Now let's show the second entry data frame"
+ "Now let's show the second entry data frame\n"
]
},
{
@@ -238,7 +245,7 @@
"id": "536ab4bb",
"metadata": {},
"source": [
- "What we really want is these three *consistent* datasets to be in the *same* single data frame. For this, we can use the `pd.concat()` function. This concatenates any given iterable of data frames."
+ "What we really want is these three _consistent_ datasets to be in the _same_ single data frame. For this, we can use the `pl.concat()` function. This concatenates any given iterable of data frames.\n"
]
},
{
@@ -248,7 +255,7 @@
"metadata": {},
"outputs": [],
"source": [
- "penguins = pd.concat(penguins_dict.values(), axis=0)\n",
+ "penguins = pl.concat(penguins_dict.values())\n",
"penguins"
]
},
@@ -263,8 +270,7 @@
"\n",
"The figure below shows such a spreadsheet: in the middle of the sheet is what looks like a data frame but there is extraneous text in cells above and below the data.\n",
"\n",
- "\n",
- "\n"
+ "\n"
]
},
{
@@ -274,8 +280,7 @@
"source": [
"This spreadsheet can be downloaded from [here](https://github.com/aeturrell/python4DS/tree/main/data) or you can load it directly from a URL. If you want to load it from your own computer's disk, you'll need to save it in a sub-folder called \"data\" first.\n",
"\n",
- "\n",
- "The top three rows and the bottom four rows are not part of the data frame. We could skip the top three rows with `skiprows`. Note that we set `skiprows=4` since the fourth row contains column names, not the data.\n"
+ "The top three rows and the bottom four rows are not part of the data frame. We could skip the top three rows by passing `read_options` to `read_excel()`. Note that we set `skip_rows=4` since the fourth row contains column names, not the data.\n"
]
},
{
@@ -285,7 +290,10 @@
"metadata": {},
"outputs": [],
"source": [
- "pd.read_excel(\"data/deaths.xlsx\", skiprows=4)"
+ "pl.read_excel(\n",
+ " \"data/deaths.xlsx\",\n",
+ " read_options={\"skip_rows\": 4},\n",
+ ")"
]
},
{
@@ -293,7 +301,7 @@
"id": "a1a8c3ca",
"metadata": {},
"source": [
- "We could also set `nrows` to omit the extraneous rows at the bottom (another option would to be to skip a set number of rows at the end using `skipfooter`)."
+ "We could also set `n_rows` inside `read_options` to omit the extraneous rows at the bottom (another option would be to skip a set number of rows at the end using `skip_footer` in `read_options`, depending on the engine).\n"
]
},
{
@@ -303,7 +311,10 @@
"metadata": {},
"outputs": [],
"source": [
- "pd.read_excel(\"data/deaths.xlsx\", skiprows=4, nrows=10)"
+ "pl.read_excel(\n",
+ " \"data/deaths.xlsx\",\n",
+ " read_options={\"skip_rows\": 4, \"n_rows\": 10},\n",
+ ")"
]
},
{
@@ -317,20 +328,20 @@
"\n",
"The underlying data in Excel spreadsheets is more complex. A cell can be one of five things:\n",
"\n",
- "- A logical, like TRUE / FALSE\n",
+ "- A logical, like TRUE / FALSE\n",
"\n",
- "- A number, like \"10\" or \"10.5\"\n",
+ "- A number, like \"10\" or \"10.5\"\n",
"\n",
- "- A date, which can also include time like \"11/1/21\" or \"11/1/21 3:00 PM\"\n",
+ "- A date, which can also include time like \"11/1/21\" or \"11/1/21 3:00 PM\"\n",
"\n",
- "- A string, like \"ten\"\n",
+ "- A string, like \"ten\"\n",
"\n",
- "- A currency, which allows numeric values in a limited range and four decimal digits of fixed precision\n",
+ "- A currency, which allows numeric values in a limited range and four decimal digits of fixed precision\n",
"\n",
"When working with spreadsheet data, it's important to keep in mind that how the underlying data is stored can be very different than what you see in the cell. For example, Excel has no notion of an integer. All numbers are stored as floating points (real number), but you can choose to display the data with a customizable number of decimal points. Similarly, dates are actually stored as numbers, specifically the number of seconds since January 1, 1970. You can customize how you display the date by applying formatting in Excel. Confusingly, it's also possible to have something that looks like a number but is actually a string (e.g. type `'10` into a cell in Excel).\n",
"\n",
- "These differences between how the underlying data are stored vs. how they're displayed can cause surprises when the data are loaded into analytical tools such as **pandas**. By default, **pandas** will guess the data type in a given column.\n",
- "A recommended workflow is to let **pandas** guess the column types initially, inspect them, and then change any data types that you want to."
+ "These differences between how the underlying data are stored vs. how they're displayed can cause surprises when the data are loaded into analytical tools such as **polars**. By default, **polars** will guess the data type in a given column.\n",
+ "A recommended workflow is to let **polars** guess the column types initially, inspect them, and then change any data types that you want to.\n"
]
},
{
@@ -340,7 +351,7 @@
"source": [
"## Writing to Excel\n",
"\n",
- "Let's create a small data frame that we can then write out. Note that `item` is a category and `quantity` is an integer."
+ "Let's create a small data frame that we can then write out. Note that `item` is a category and `quantity` is an integer.\n"
]
},
{
@@ -350,8 +361,11 @@
"metadata": {},
"outputs": [],
"source": [
- "bake_sale = pd.DataFrame(\n",
- " {\"item\": pd.Categorical([\"brownie\", \"cupcake\", \"cookie\"]), \"quantity\": [10, 5, 8]}\n",
+ "bake_sale = pl.DataFrame(\n",
+ " {\n",
+ " \"item\": pl.Series([\"brownie\", \"cupcake\", \"cookie\"], dtype=pl.Categorical),\n",
+ " \"quantity\": [10, 5, 8],\n",
+ " }\n",
")\n",
"bake_sale"
]
@@ -361,17 +375,17 @@
"id": "345bca3d",
"metadata": {},
"source": [
- "You can write data back to disk as an Excel file using the `.to_excel()` function. The `index=False` keyword argument just writes the two columns without the index that was automatically added in the last step."
+ "You can write data back to disk as an Excel file using the `.write_excel()` method. Polars does not use a row index like pandas, so only the columns in the DataFrame are written by default.\n"
]
},
{
- "cell_type": "markdown",
+ "cell_type": "code",
+ "execution_count": null,
"id": "1fc17141",
"metadata": {},
+ "outputs": [],
"source": [
- "```python\n",
- "bake_sale.to_excel(\"data/bake_sale.xlsx\", index=False)\n",
- "```"
+ "bake_sale.write_excel(\"data/bake_sale.xlsx\")"
]
},
{
@@ -381,7 +395,7 @@
"source": [
"The figure below shows what the data looks like in Excel.\n",
"\n",
- ""
+ "\n"
]
},
{
@@ -389,7 +403,7 @@
"id": "8d555c84",
"metadata": {},
"source": [
- "Just like reading from a CSV, information on data type is lost when we read the data back in—you can see this is you read the data back in and check the `info` for the data types. Although we kept `int64` because **pandas** recognise that the second column was of integer type, we lost the categorical data type for \"item\". This data type loss makes Excel files unreliable for caching interim results."
+ "Just like reading from a CSV, information on data type is lost when we read the data back in—you can see this if you read the data back in and check the `schema` for the data types. Although we kept `Int64` because **polars** recognised that the second column was of integer type, we lost the categorical data type for \"item\". This data type loss makes Excel files unreliable for caching interim results.\n"
]
},
{
@@ -399,7 +413,7 @@
"metadata": {},
"outputs": [],
"source": [
- "pd.read_excel(\"data/bake_sale.xlsx\").info()"
+ "pl.read_excel(\"data/bake_sale.xlsx\").schema"
]
},
{
@@ -409,14 +423,11 @@
"source": [
"### Formatted Output\n",
"\n",
- "If you need more formatting options and more control over how you write spreadsheets, check out the documentation for [openpyxl](https://openpyxl.readthedocs.io/) which can do pretty much everything you imagine. Generally, releasing data in spreadsheets is not the best option: but if you do want to release data in spreadsheets according to best practice, then check out [gptables](https://gptables.readthedocs.io/)."
+ "If you need more formatting options and more control over how you write spreadsheets, check out the documentation for [openpyxl](https://openpyxl.readthedocs.io/) which can do pretty much everything you imagine. Generally, releasing data in spreadsheets is not the best option: but if you do want to release data in spreadsheets according to best practice, then check out [gptables](https://gptables.readthedocs.io/).\n"
]
}
],
"metadata": {
- "interpreter": {
- "hash": "9d7534ecd9fbc7d385378f8400cf4d6cb9c6175408a574f1c99c5269f08771cc"
- },
"jupytext": {
"cell_metadata_filter": "-all",
"encoding": "# -*- coding: utf-8 -*-",
@@ -424,7 +435,7 @@
"main_language": "python"
},
"kernelspec": {
- "display_name": "Python 3 (ipykernel)",
+ "display_name": "python4ds",
"language": "python",
"name": "python3"
},
diff --git a/uv.lock b/uv.lock
index 8316606..8e7d9ad 100644
--- a/uv.lock
+++ b/uv.lock
@@ -354,6 +354,19 @@ wheels = [
{ url = "https://files.pythonhosted.org/packages/b5/fd/afcd0496feca3276f509df3dbd5dae726fcc756f1a08d9e25abe1733f962/executing-2.1.0-py2.py3-none-any.whl", hash = "sha256:8d63781349375b5ebccc3142f4b30350c0cd9c79f921cde38be2be4637e98eaf", size = 25805, upload-time = "2024-09-01T12:37:33.007Z" },
]
+[[package]]
+name = "fastexcel"
+version = "0.19.0"
+source = { registry = "https://pypi.org/simple" }
+sdist = { url = "https://files.pythonhosted.org/packages/0d/c8/3b09911348e9c64dbf41096d3e8f0e93c141a23990ec9f32514111bd5f55/fastexcel-0.19.0.tar.gz", hash = "sha256:216c3719ee90963bd93a0bf8c10b177233046ac975b67651152fdaedd3c99aa1", size = 60323, upload-time = "2026-01-20T11:17:37.253Z" }
+wheels = [
+ { url = "https://files.pythonhosted.org/packages/d1/e0/3820e93ea606549cfddb8c437141dd69f2b245e74785efc8bd7511ba909d/fastexcel-0.19.0-cp310-abi3-macosx_10_12_x86_64.whl", hash = "sha256:68601072a0b4b4277c165b68f1055f88ef7ffe7ed6f08c1eeda0f0271e3f7da0", size = 3082362, upload-time = "2026-01-20T11:17:27.157Z" },
+ { url = "https://files.pythonhosted.org/packages/66/0f/b42dc09515879192919942157292912393584045fd8bad98bd92961d4c30/fastexcel-0.19.0-cp310-abi3-macosx_11_0_arm64.whl", hash = "sha256:c8a87d94445678e7e3f46a6aa39d2afaee5b88a983ec3661143a6488d8955f44", size = 2864365, upload-time = "2026-01-20T11:17:28.786Z" },
+ { url = "https://files.pythonhosted.org/packages/8e/4a/bc358b20fcff64b4c14ff7d7a0e1f797792b8b77e30ae755873c02362538/fastexcel-0.19.0-cp310-abi3-manylinux_2_17_aarch64.manylinux2014_aarch64.whl", hash = "sha256:e94fc1be6642555f277af792c22a9f80ec9b4d640d9690f00abb822b6d865069", size = 3186426, upload-time = "2026-01-20T11:17:19.087Z" },
+ { url = "https://files.pythonhosted.org/packages/58/ae/d2ffdc5ad14190153e2422fc90a1052a4b0c3086d24cb8ae8967575321d8/fastexcel-0.19.0-cp310-abi3-manylinux_2_17_x86_64.manylinux2014_x86_64.whl", hash = "sha256:334f9f40cd68b5924a712b6c104949757a0b8ad8a7e3fa3f3fad1c1ebc00258b", size = 3365628, upload-time = "2026-01-20T11:17:21.116Z" },
+ { url = "https://files.pythonhosted.org/packages/6e/67/5f6d4e7760dc3dd8244cd124dabdd5bb7622bf1197edcc2513648847690e/fastexcel-0.19.0-cp310-abi3-win_amd64.whl", hash = "sha256:fbbdf9de79c3ef3572809bb187927c0dc5840968ffe513ea015a383024b7c6b0", size = 2905173, upload-time = "2026-01-20T11:17:33.687Z" },
+]
+
[[package]]
name = "fastjsonschema"
version = "2.21.1"
@@ -1725,10 +1738,11 @@ wheels = [
[[package]]
name = "python4ds"
-version = "1.0.4"
+version = "0.0.1"
source = { virtual = "." }
dependencies = [
{ name = "beautifulsoup4" },
+ { name = "fastexcel" },
{ name = "graphviz" },
{ name = "ibis-framework", extra = ["sqlite"] },
{ name = "ipykernel" },
@@ -1759,12 +1773,14 @@ dependencies = [
{ name = "toml" },
{ name = "watermark" },
{ name = "wbgapi" },
+ { name = "xlsxwriter" },
{ name = "yfinance" },
]
[package.metadata]
requires-dist = [
{ name = "beautifulsoup4", specifier = ">=4.12.3" },
+ { name = "fastexcel", specifier = ">=0.19.0" },
{ name = "graphviz", specifier = ">=0.20.3" },
{ name = "ibis-framework", extras = ["sqlite"], specifier = ">=9.5.0" },
{ name = "ipykernel", specifier = ">=6.29.5" },
@@ -1795,6 +1811,7 @@ requires-dist = [
{ name = "toml", specifier = ">=0.10.2" },
{ name = "watermark", specifier = ">=2.5.0" },
{ name = "wbgapi", specifier = ">=1.0.14" },
+ { name = "xlsxwriter", specifier = ">=3.2.0" },
{ name = "yfinance", specifier = ">=1.2.1" },
]
@@ -2463,6 +2480,15 @@ wheels = [
{ url = "https://files.pythonhosted.org/packages/21/02/88b65cc394961a60c43c70517066b6b679738caf78506a5da7b88ffcb643/widgetsnbextension-4.0.13-py3-none-any.whl", hash = "sha256:74b2692e8500525cc38c2b877236ba51d34541e6385eeed5aec15a70f88a6c71", size = 2335872, upload-time = "2024-08-22T12:18:19.491Z" },
]
+[[package]]
+name = "xlsxwriter"
+version = "3.2.9"
+source = { registry = "https://pypi.org/simple" }
+sdist = { url = "https://files.pythonhosted.org/packages/46/2c/c06ef49dc36e7954e55b802a8b231770d286a9758b3d936bd1e04ce5ba88/xlsxwriter-3.2.9.tar.gz", hash = "sha256:254b1c37a368c444eac6e2f867405cc9e461b0ed97a3233b2ac1e574efb4140c", size = 215940, upload-time = "2025-09-16T00:16:21.63Z" }
+wheels = [
+ { url = "https://files.pythonhosted.org/packages/3a/0c/3662f4a66880196a590b202f0db82d919dd2f89e99a27fadef91c4a33d41/xlsxwriter-3.2.9-py3-none-any.whl", hash = "sha256:9a5db42bc5dff014806c58a20b9eae7322a134abb6fce3c92c181bfb275ec5b3", size = 175315, upload-time = "2025-09-16T00:16:20.108Z" },
+]
+
[[package]]
name = "yfinance"
version = "1.2.1"
diff --git a/visualise.quarto_ipynb_1 b/visualise.quarto_ipynb_1
new file mode 100644
index 0000000..2ef6d2f
--- /dev/null
+++ b/visualise.quarto_ipynb_1
@@ -0,0 +1,136 @@
+{
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Visualisation {#sec-visualise}\n",
+ "\n",
+ "After reading the first part of the book, you understand the basics of the most important tools for doing data science. Now it’s time to start diving into the details. In this part of the book, you’ll learn about visualising data in further depth (in @sec-vis-layers), and get further stuck into the details of the different kinds of data visualisation (in @sec-exploratory-data-analysis and @sec-communicate-plots). In this short chapter, we discuss the different ways to create visualisations, and the different purposes of visualisations.\n",
+ "\n",
+ "## Philosophies of data visualisation\n",
+ "\n",
+ "There are broadly two categories of approach to using code to create data visualisations: *imperative* (build what you want from individual elements) and *declarative* (say what you want from a list of pre-existing options). Choosing which to use involves a trade-off: imperative libraries offer you flexibility but at the cost of some verbosity; declarative libraries offer you a quick way to plot your data, but only if it’s in the right format to begin with, and customisation to special chart types is more difficult.\n",
+ "\n",
+ "Python has many excellent plotting packages, including perhaps the most powerful imperative plotting package around, **matplotlib**, and an amazing declarative library that we already saw, **lets-plot**. These two libraries will get you a long way, and each could be worthy of an entire book themselves. Fortunately for us, though, we can do 95% of what we need with a small number of commands from one or the other of them. In general, to keep this book as light as possible, we've opted to use **lets-plot** wherever possible—and @sec-vis-layers is going to take you on a more in-depth tour of how to use it yourself.\n",
+ "\n",
+ "## Purposes of data visualisation\n",
+ "\n",
+ "Data visualisation has all kinds of different purposes. It can be useful to bear in mind three broad categories of visualisation that are out there:\n",
+ "\n",
+ "- exploratory\n",
+ "- scientific\n",
+ "- narrative\n",
+ "\n",
+ "Let's look at each in a bit more detail.\n",
+ "\n",
+ "### Exploratory Data Viz\n",
+ "\n",
+ "The first of the three kinds is *exploratory data visualisation*, and it's the kind that you do when you're looking and data and trying to understand it. Just plotting the data is a really good strategy for getting a feel for any issues there might be. This is perhaps most famously demonstrated by Anscombe's quartet: four different datasets with the same mean, standard deviation, and correlation but very different data distributions."
+ ],
+ "id": "f3331573"
+ },
+ {
+ "cell_type": "code",
+ "metadata": {},
+ "source": [
+ "#| echo: false\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "import matplotlib.pyplot as plt\n",
+ "import matplotlib_inline.backend_inline\n",
+ "\n",
+ "# Plot settings\n",
+ "plt.style.use(\"https://github.com/aeturrell/python4DS/raw/main/plot_style.txt\")\n",
+ "matplotlib_inline.backend_inline.set_matplotlib_formats(\"svg\")\n",
+ "\n",
+ "# Set max rows displayed for readability\n",
+ "pd.set_option(\"display.max_rows\", 6)\n",
+ "\n",
+ "x = [10, 8, 13, 9, 11, 14, 6, 4, 12, 7, 5]\n",
+ "y1 = [8.04, 6.95, 7.58, 8.81, 8.33, 9.96, 7.24, 4.26, 10.84, 4.82, 5.68]\n",
+ "y2 = [9.14, 8.14, 8.74, 8.77, 9.26, 8.10, 6.13, 3.10, 9.13, 7.26, 4.74]\n",
+ "y3 = [7.46, 6.77, 12.74, 7.11, 7.81, 8.84, 6.08, 5.39, 8.15, 6.42, 5.73]\n",
+ "x4 = [8, 8, 8, 8, 8, 8, 8, 19, 8, 8, 8]\n",
+ "y4 = [6.58, 5.76, 7.71, 8.84, 8.47, 7.04, 5.25, 12.50, 5.56, 7.91, 6.89]\n",
+ "\n",
+ "datasets = {\"I\": (x, y1), \"II\": (x, y2), \"III\": (x, y3), \"IV\": (x4, y4)}\n",
+ "\n",
+ "fig, axs = plt.subplots(\n",
+ " 2,\n",
+ " 2,\n",
+ " sharex=True,\n",
+ " sharey=True,\n",
+ " figsize=(10, 6),\n",
+ " gridspec_kw={\"wspace\": 0.08, \"hspace\": 0.08},\n",
+ ")\n",
+ "axs[0, 0].set(xlim=(0, 20), ylim=(2, 14))\n",
+ "axs[0, 0].set(xticks=(0, 10, 20), yticks=(4, 8, 12))\n",
+ "\n",
+ "for ax, (label, (x, y)) in zip(axs.flat, datasets.items()):\n",
+ " ax.text(0.1, 0.9, label, fontsize=20, transform=ax.transAxes, va=\"top\")\n",
+ " ax.tick_params(direction=\"in\", top=True, right=True)\n",
+ " ax.plot(x, y, \"o\")\n",
+ "\n",
+ " # linear regression\n",
+ " p1, p0 = np.polyfit(x, y, deg=1) # slope, intercept\n",
+ " ax.axline(xy1=(0, p0), slope=p1, color=\"r\", lw=2)\n",
+ "\n",
+ " # add text box for the statistics\n",
+ " stats = (\n",
+ " f\"$\\\\mu$ = {np.mean(y):.2f}\\n\"\n",
+ " f\"$\\\\sigma$ = {np.std(y):.2f}\\n\"\n",
+ " f\"$r$ = {np.corrcoef(x, y)[0][1]:.2f}\"\n",
+ " )\n",
+ " bbox = dict(boxstyle=\"round\", fc=\"blanchedalmond\", ec=\"orange\", alpha=0.5)\n",
+ " ax.text(\n",
+ " 0.95,\n",
+ " 0.07,\n",
+ " stats,\n",
+ " fontsize=9,\n",
+ " bbox=bbox,\n",
+ " transform=ax.transAxes,\n",
+ " horizontalalignment=\"right\",\n",
+ " )\n",
+ "\n",
+ "plt.suptitle(\"Anscombe's Quartet\")\n",
+ "plt.show()"
+ ],
+ "id": "64a0e7f6",
+ "execution_count": null,
+ "outputs": []
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Exploratory visualisation is usually quick and dirty, and flexible too. Some exploratory data viz can be automated, and there's a whole host of packages to help with this, including [**skimpy**](https://aeturrell.github.io/skimpy/).\n",
+ "\n",
+ "Beyond you and perhaps your co-authors/collaborators, however, not many other people should be seeing your exploratory visualisation! They will typically be worked up quickly, be numerous, and be throw-away. We'll look more at this in @sec-exploratory-data-analysis.\n",
+ "\n",
+ "### Scientific Data Viz\n",
+ "\n",
+ "The second kind, scientific data visualisation, is the prime cut of your exploratory visualisation. It's the kind of plot you might include in a more technical paper, the picture that says a thousand words. I often think of the first image of a black hole @akiyama2019first as a prime example of this. You can get away with having a high density of information in a scientific plot and, in short format journals, you may need to. The journal Physical Review Letters, which has an 8 page limit, has a classic of this genre in more or less every issue. Ensuring that important values can be accurately read from the plot is especially important in these kinds of charts. But they can also be the kind of plot that presents the killer results in a study; they might not be exciting to people who don't look at charts for a living, but they might be exciting and, just as importantly, understandable by your peers.\n",
+ "\n",
+ "This type of visualisation is especially popular in the big science journals like *Nature* and *Science*, where space is at a premium. We won't cover this type of plot in this book, because it tends to be very bespoke.\n",
+ "\n",
+ "### Narrative Data Viz\n",
+ "\n",
+ "The third and final kind is narrative data visualisation. This is the one that requires the most thought in the step where you go from the first view to the end product. It's a visualisation that doesn't just show a picture, but gives an insight. These are the kind of visualisations that you might see in the *Financial Times*, *The Economist*, or on the *BBC News* website. They come with aids that help the viewer focus on the aspects that the creator wanted them to (you can think of these aids or focuses as doing for visualisation what bold font does for text). They're well worth using in your work, especially if you're trying to communicate a particular narrative, and especially if the people you're communicating with don't have deep knowledge of the topic. You might use them in a paper that you hope will have a wide readership, in a blog post summarising your work, or in a report intended for a policymaker.\n",
+ "\n",
+ "You can find more information on the topic of communicating via data visualisations in the @sec-communicate-plots chapter."
+ ],
+ "id": "30b9ff30"
+ }
+ ],
+ "metadata": {
+ "kernelspec": {
+ "name": "python3",
+ "language": "python",
+ "display_name": "Python 3 (ipykernel)",
+ "path": "/Users/omagic/Documents/GitHub/python4DSpolars/.venv/share/jupyter/kernels/python3"
+ }
+ },
+ "nbformat": 4,
+ "nbformat_minor": 5
+}
\ No newline at end of file
diff --git a/webscraping-and-apis.ipynb b/webscraping-and-apis.ipynb
index 9366eb6..5171a4d 100644
--- a/webscraping-and-apis.ipynb
+++ b/webscraping-and-apis.ipynb
@@ -10,7 +10,7 @@
"\n",
"## Introduction\n",
"\n",
- "This chapter will show you how to work with online data that is either obtained from webpages via webscraping or more directly over the internet via an API. An important principle is always to use an API if one is available as this is designed to pass information directly into your Python session and will save you a lot of effort."
+ "This chapter will show you how to work with online data that is either obtained from webpages via webscraping or more directly over the internet via an API. An important principle is always to use an API if one is available as this is designed to pass information directly into your Python session and will save you a lot of effort.\n"
]
},
{
@@ -56,7 +56,7 @@
"\n",
"As a brief example, in the US, lists of ingredients and instructions are not copyrightable, so copyright can not be used to protect a recipe. But if that list of recipes is accompanied by substantial novel literary content, that is copyrightable. This is why when you’re looking for a recipe on the internet there’s always so much content beforehand.\n",
"\n",
- "If you do need to scrape original content (like text or images), you may still be protected under the doctrine of fair use. Fair use is not a hard and fast rule, but weighs up a number of factors. It’s more likely to apply if you are collecting the data for research or non-commercial purposes and if you limit what you scrape to just what you need."
+ "If you do need to scrape original content (like text or images), you may still be protected under the doctrine of fair use. Fair use is not a hard and fast rule, but weighs up a number of factors. It’s more likely to apply if you are collecting the data for research or non-commercial purposes and if you limit what you scrape to just what you need.\n"
]
},
{
@@ -67,9 +67,9 @@
"source": [
"### Prerequisites\n",
"\n",
- "You will need to install the **pandas** package for this chapter. We'll use **seaborn** too, which you should already have installed. You will also need to install the **beautifulsoup**, **pandas-datareader**, and **wbgapi** packages in your terminal using `uv add beautifulsoup4`, `uv add pandas-datareader`, and `uv add wbgapi` respectively. We'll also use two built-in packages, **textwrap** and **requests**.\n",
+ "You will need to install the **pandas** and **polars** package for this chapter. We'll use **seaborn** too, which you should already have installed. You will also need to install the **beautifulsoup**, **pandas-datareader**, and **wbgapi** packages in your terminal using `uv add beautifulsoup4`, and `uv add wbgapi` respectively. We'll also use two built-in packages, **textwrap** and **requests**.\n",
"\n",
- "To kick off, let's import some of the packages we need (it's always good practice to import the packages you need at the top of a script or notebook)."
+ "To kick off, let's import some of the packages we need (it's always good practice to import the packages you need at the top of a script or notebook).\n"
]
},
{
@@ -81,12 +81,11 @@
"source": [
"import textwrap\n",
"\n",
+ "import lets_plot as lp\n",
"import pandas as pd\n",
+ "import polars as pl\n",
"import requests\n",
- "from bs4 import BeautifulSoup\n",
- "from lets_plot import *\n",
- "\n",
- "LetsPlot.setup_html()"
+ "from bs4 import BeautifulSoup"
]
},
{
@@ -95,9 +94,9 @@
"id": "f43a5237",
"metadata": {},
"source": [
- "## Extracting Data from Files on the Internet using **pandas**\n",
+ "## Extracting Data from Files on the Internet using **polars**\n",
"\n",
- "It's easy to read data from the internet once you have the url and file type. Here, for instance, is an example that reads in the 'storms' dataset, which is stored as a CSV file in a URL (we'll only grab the first 10 rows):"
+ "It's easy to read data from the internet once you have the url and file type. Here, for instance, is an example that reads in the 'storms' dataset, which is stored as a CSV file in a URL (we'll only grab the first 10 rows):\n"
]
},
{
@@ -107,8 +106,8 @@
"metadata": {},
"outputs": [],
"source": [
- "pd.read_csv(\n",
- " \"https://vincentarelbundock.github.io/Rdatasets/csv/dplyr/storms.csv\", nrows=10\n",
+ "pl.read_csv(\n",
+ " \"https://vincentarelbundock.github.io/Rdatasets/csv/dplyr/storms.csv\", n_rows=10\n",
")"
]
},
@@ -122,7 +121,7 @@
"\n",
"Using an API (application programming interface) is another way to draw down information from the interweb. Their just a way for one tool, say Python, to speak to another tool, say a server, and usefully exchange information. The classic use case would be to post a request for data that fits a certain query via an API and to get a download of that data back in return. (You should always preferentially use an API over webscraping a site.)\n",
"\n",
- "Because they are designed to work with any tool, you don't actually need a programming language to interact with an API, it's just a *lot* easier if you do.\n",
+ "Because they are designed to work with any tool, you don't actually need a programming language to interact with an API, it's just a _lot_ easier if you do.\n",
"\n",
"::: {.callout-note}\n",
"An API key is needed in order to access some APIs. Sometimes all you need to do is register with site, in other cases you may have to pay for access.\n",
@@ -132,13 +131,13 @@
"\n",
"An API has an 'endpoint', the base url, and then a URL that encodes the question. Let's see an example with the ONS API for which the endpoint is \"https://api.beta.ons.gov.uk/v1/\". The rest of the API has the form 'data?uri=' and then the long ID of both the timeseries (jp9z) and then the dataset (LMS), which is vacancies in the UK services sector.\n",
"\n",
- "The data that are returned by APIs are typically in JSON format, which looks a lot like a nested Python dictionary and its entries can be accessed in the same way--this is what is happening when getting the series' title in the example below. JSON is not good for analysis, so we'll use **pandas** to put the data into shape."
+ "The data that are returned by APIs are typically in JSON format, which looks a lot like a nested Python dictionary and its entries can be accessed in the same way--this is what is happening when getting the series' title in the example below. JSON is not good for analysis, so we'll use **polars** to put the data into shape.\n"
]
},
{
"cell_type": "code",
"execution_count": null,
- "id": "c4226d67",
+ "id": "6107093c",
"metadata": {},
"outputs": [],
"source": [
@@ -147,18 +146,36 @@
"# Get the data from the ONS API:\n",
"json_data = requests.get(url).json()\n",
"\n",
- "# Prep the data for a quick plot\n",
"title = json_data[\"description\"][\"title\"]\n",
+ "\n",
+ "# Convert dates using string operations\n",
"df = (\n",
- " pd.DataFrame(pd.json_normalize(json_data[\"months\"]))\n",
- " .assign(\n",
- " date=lambda x: pd.to_datetime(x[\"date\"]),\n",
- " value=lambda x: pd.to_numeric(x[\"value\"]),\n",
+ " pl.DataFrame(json_data[\"months\"])\n",
+ " .with_columns(\n",
+ " [\n",
+ " # Add day to make it a valid date string\n",
+ " (pl.col(\"date\") + \"-01\").str.to_date(format=\"%Y %b-%d\").alias(\"date\"),\n",
+ " pl.col(\"value\").cast(pl.Float64).alias(\"value\"),\n",
+ " ]\n",
" )\n",
- " .set_index(\"date\")\n",
+ " .drop_nulls(\"date\")\n",
+ " .sort(\"date\")\n",
+ ")\n",
+ "\n",
+ "\n",
+ "# Initialize the library\n",
+ "lp.LetsPlot.setup_html()\n",
+ "\n",
+ "# Create plot using the alias\n",
+ "chart = (\n",
+ " lp.ggplot(df, lp.aes(x=\"date\", y=\"value\"))\n",
+ " + lp.geom_line(size=2.0, color=\"steelblue\")\n",
+ " + lp.ggtitle(title)\n",
+ " + lp.ylim(0, df[\"value\"].max() * 1.2)\n",
+ " + lp.theme_classic()\n",
")\n",
"\n",
- "df[\"value\"].plot(title=title, ylim=(0, df[\"value\"].max() * 1.2), lw=3.0);"
+ "chart"
]
},
{
@@ -167,37 +184,9 @@
"id": "670ce0bb",
"metadata": {},
"source": [
- "We've talked about *reading* APIs. You can also create your own to serve up data, models, whatever you like! This is an advanced topic and we won't cover it; but if you do need to, the simplest way is to use [Fast API](https://fastapi.tiangolo.com/). You can find some short video tutorials for Fast API [here](https://calmcode.io/fastapi/hello-world.html).\n",
- "\n",
- "### Pandas Datareader: an easier way to interact with (some) APIs\n",
- "\n",
- "Although it didn't take much code to get the ONS data, it would be even better if it was just a single line, wouldn't it? Fortunately there are some packages out there that make this easy, but it does depend on the API (and APIs come and go over time).\n",
- "\n",
- "By far the most comprehensive library for accessing extra APIs is [**pandas-datareader**](https://pandas-datareader.readthedocs.io/en/latest/), which provides convenient access to:\n",
- "\n",
- "- FRED\n",
- "- Quandl\n",
- "- World Bank\n",
- "- OECD\n",
- "- Eurostat\n",
+ "We've talked about _reading_ APIs. You can also create your own to serve up data, models, whatever you like! This is an advanced topic and we won't cover it; but if you do need to, the simplest way is to use [Fast API](https://fastapi.tiangolo.com/). You can find some short video tutorials for Fast API [here](https://calmcode.io/fastapi/hello-world.html).\n",
"\n",
- "and more.\n",
- "\n",
- "Let's see an example using FRED (the Federal Reserve Bank of St. Louis' economic data library). This time, let's look at the UK unemployment rate:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "id": "bf758fb4",
- "metadata": {},
- "outputs": [],
- "source": [
- "import pandas_datareader.data as web\n",
- "\n",
- "df_u = web.DataReader(\"LRHUTTTTGBM156S\", \"fred\")\n",
- "\n",
- "df_u.plot(title=\"UK unemployment (percent)\", legend=False, ylim=(2, 6), lw=3.0);"
+ "### Accessing World Bank Data with wbgapi\n"
]
},
{
@@ -206,7 +195,9 @@
"id": "0613aefb",
"metadata": {},
"source": [
- "And, because it's also a really useful one, let's see how to use the [**wbgapi**](https://pypi.org/project/wbgapi/) package to access World Bank data. (**pandas-datareader** used to provide a World Bank reader too, but it has not been actively maintained, so we prefer **wbgapi** for new work.)"
+ "While APIs can be accessed directly using tools like requests, some specialized libraries make working with structured datasets much easier. One such example is wbgapi, which provides a convenient interface for accessing World Bank data.\n",
+ "\n",
+ "Let’s look at an example using World Bank data on CO₂-equivalent emissions per capita:\n"
]
},
{
@@ -224,18 +215,21 @@
"import wbgapi as wb\n",
"\n",
"indicator_code = \"EN.GHG.ALL.PC.CE.AR5\"\n",
+ "\n",
"df = (\n",
- " wb.data.DataFrame(\n",
- " indicator_code,\n",
- " [\"USA\", \"CHN\", \"IND\", \"EAS\", \"ECS\"], # country and region codes\n",
- " time=range(2019, 2020),\n",
- " labels=True,\n",
+ " pl.from_pandas(\n",
+ " wb.data.DataFrame(\n",
+ " indicator_code,\n",
+ " [\"USA\", \"CHN\", \"IND\", \"EAS\", \"ECS\"],\n",
+ " time=range(2019, 2020),\n",
+ " labels=True,\n",
+ " ).reset_index()\n",
" )\n",
- " .rename(columns={\"Country\": \"country\", \"YR2019\": indicator_code})\n",
- " .reset_index(drop=True)\n",
+ " .rename({\"Country\": \"country\"})\n",
+ " .with_columns(pl.col(\"country\").map_elements(lambda x: textwrap.fill(x, 10)))\n",
+ " .sort(indicator_code, descending=True)\n",
")\n",
- "df[\"country\"] = df[\"country\"].apply(lambda x: textwrap.fill(x, 10)) # wrap long names\n",
- "df = df.sort_values(indicator_code) # re-order\n",
+ "\n",
"df.head()"
]
},
@@ -246,19 +240,26 @@
"metadata": {},
"outputs": [],
"source": [
- "(\n",
- " ggplot(df, aes(x=\"country\", y=indicator_code))\n",
- " + geom_bar(aes(fill=\"country\"), color=\"black\", alpha=0.8, stat=\"identity\")\n",
- " + scale_fill_discrete()\n",
- " + theme_minimal()\n",
- " + theme(legend_position=\"none\")\n",
- " + ggsize(600, 400)\n",
- " + labs(\n",
+ "lp.LetsPlot.setup_html()\n",
+ "\n",
+ "country_order = df[\"country\"].to_list()\n",
+ "\n",
+ "plot = (\n",
+ " lp.ggplot(df, lp.aes(x=\"country\", y=indicator_code))\n",
+ " + lp.geom_bar(lp.aes(fill=\"country\"), color=\"black\", alpha=0.8, stat=\"identity\")\n",
+ " + lp.scale_x_discrete(limits=country_order)\n",
+ " + lp.scale_fill_discrete()\n",
+ " + lp.theme_minimal()\n",
+ " + lp.theme(legend_position=\"none\")\n",
+ " + lp.ggsize(600, 400)\n",
+ " + lp.labs(\n",
" subtitle=\"Greenhouse gases (CO2-equivalent metric tons per capita, 2019)\",\n",
" title=\"The USA leads the world on per-capita emissions\",\n",
" y=\"\",\n",
" )\n",
- ")"
+ ")\n",
+ "\n",
+ "plot.show()"
]
},
{
@@ -267,15 +268,19 @@
"id": "b7bf16d7",
"metadata": {},
"source": [
- "### The OECD API\n",
+ "### The Eurostat SDMX API\n",
+ "\n",
+ "Sometimes it’s convenient to use APIs directly. The Eurostat API provides access to a massive repository of European statistical data using the SDMX (Statistical Data and Metadata eXchange) standard. While Eurostat offers multiple formats, using the SDMX-ML (XML) format via the sdmx1 library allows us to pull structured data into the Python ecosystem with high precision.\n",
+ "\n",
+ "Key to using the Eurostat API is understanding the Data Structure Definition (DSD). Every dataset is essentially a multidimensional \"cube\" where each dimension (like Geography, Unit, or Frequency) has specific codes.\n",
"\n",
- "Sometimes it's convenient to use APIs directly, and, as an example, the OECD API comes with a LOT of complexity that direct access can take advantage of. The OECD API makes data available in both JSON and XML formats, and we'll use [**pandasdmx**](https://pandasdmx.readthedocs.io/) (aka the Statistical Data and Metadata eXchange (SDMX) package for the Python data ecosystem) to pull down the XML format data and turn it into a regular **pandas** data frame.\n",
+ "To find the exact codes you need:\n",
"\n",
- "Now, key to using the OECD API is knowledge of its many codes: for countries, times, resources, and series. You can find some broad guidance on what codes the API uses [here](https://data.oecd.org/api/sdmx-ml-documentation/) but to find exactly what you need can be a bit tricky. Two tips are:\n",
- "1. If you know what you're looking for is in a particular named dataset, eg \"QNA\" (Quarterly National Accounts), put `https://stats.oecd.org/restsdmx/sdmx.ashx/GetDataStructure/QNA/all?format=SDMX-ML` into your browser and look through the XML file; you can pick out the sub-codes and the countries that are available.\n",
- "2. Browse around on https://stats.oecd.org/ and use Customise then check all the \"Use Codes\" boxes to see whatever your browsing's code names.\n",
+ "The Data Browser: Browse the Eurostat Data Navigation Tree. Once you find a table (e.g., \"HICP - monthly data\"), the \"Dataset Code\" (like prc_hicp_manr) is shown in brackets.\n",
"\n",
- "Let's see an example of this in action. We'd like to see the productivity (GDP per hour) data for a range of countries since 2010. We are going to be in the productivity resource (code \"PDB_LV\") and we want the USD current prices (code \"CPC\") measure of GDP per employed worker (code \"T_GDPEMP) from 2010 onwards (code \"startTime=2010\"). We'll grab this for some developed countries where productivity measurements might be slightly more comparable. The comments below explain what's happening in each step."
+ "Positional Keys: Eurostat's REST API expects a \"key string\" where codes are placed in a specific order separated by dots (e.g., Freq.Unit.Item.Geo). If you know the order, you can \"slice\" the data cube directly.\n",
+ "\n",
+ "Let’s see an example of this in action. We want to see the Harmonised Index of Consumer Prices (HICP)—specifically the annual rate of change for all items—for Germany and France. We will use the resource prc_hicp_manr, requesting Monthly frequency (M), the Annual Rate of Change unit (RCH_A), and the \"All-items\" classification (CP00).\n"
]
},
{
@@ -285,18 +290,33 @@
"metadata": {},
"source": [
"```python\n",
- "import pandasdmx as pdmx\n",
- "# Tell pdmx we want OECD data\n",
- "oecd = pdmx.Request(\"OECD\")\n",
- "# Set out everything about the request in the format specified by the OECD API\n",
- "data = oecd.data(\n",
- " resource_id=\"PDB_LV\",\n",
- " key=\"GBR+FRA+CAN+ITA+DEU+JPN+USA.T_GDPEMP.CPC/all?startTime=2010\",\n",
- ").to_pandas()\n",
+ "import sdmx\n",
+ "import polars as pl\n",
+ "\n",
+ "# Tell sdmx we want ESTAT data\n",
+ "client = sdmx.Client('ESTAT')\n",
+ "\n",
+ "# 2. Build the URL-style positional key\n",
+ "# Format: [Freq].[Unit].[Coicop].[Geo]\n",
+ "# We use '+' to join multiple countries (DE and FR)\n",
+ "resource_id = 'prc_hicp_manr'\n",
+ "key_string = 'M.RCH_A.CP00.DE+FR'\n",
+ "\n",
+ "# 3. Fetch the data directly\n",
+ "# 'startPeriod' limits the timeline to recent data\n",
+ "response = client.data(\n",
+ " resource_id=resource_id,\n",
+ " key=key_string,\n",
+ " params={'startPeriod': '2024-01'}\n",
+ ")\n",
"\n",
- "df = pd.DataFrame(data).reset_index()\n",
- "df.head()\n",
- "```"
+ "# 4. Convert the SDMX-ML response to a Polars DataFrame\n",
+ "# We bridge through Pandas as sdmx1 is optimized for it\n",
+ "df_pd = sdmx.to_pandas(response).to_frame(name='value').reset_index()\n",
+ "df = pl.from_pandas(df_pd)\n",
+ "\n",
+ "print(df.head())\n",
+ "```\n"
]
},
{
@@ -305,13 +325,13 @@
"id": "e5cac233",
"metadata": {},
"source": [
- "| | LOCATION | SUBJECT | MEASURE | TIME_PERIOD | value |\n",
- "|--:|---------:|---------:|--------:|------------:|-------------:|\n",
- "| 0 | CAN | T_GDPEMP | CPC | 2010 | 78848.604088 |\n",
- "| 1 | CAN | T_GDPEMP | CPC | 2011 | 81422.364748 |\n",
- "| 2 | CAN | T_GDPEMP | CPC | 2012 | 82663.028058 |\n",
- "| 3 | CAN | T_GDPEMP | CPC | 2013 | 86368.582158 |\n",
- "| 4 | CAN | T_GDPEMP | CPC | 2014 | 89617.632446 |"
+ "| | TIME_PERIOD | geo | unit | freq | coicop | value |\n",
+ "| --: | ----------: | :-- | :---- | :--- | :----- | ----: |\n",
+ "| 0 | 2024-01 | DE | RCH_A | M | CP00 | 3.1 |\n",
+ "| 1 | 2024-02 | DE | RCH_A | M | CP00 | 2.7 |\n",
+ "| 2 | 2024-03 | DE | RCH_A | M | CP00 | 2.3 |\n",
+ "| 3 | 2024-04 | DE | RCH_A | M | CP00 | 2.4 |\n",
+ "| 4 | 2024-05 | DE | RCH_A | M | CP00 | 2.8 |\n"
]
},
{
@@ -320,7 +340,7 @@
"id": "302326b4",
"metadata": {},
"source": [
- "Great that worked! We have data in a nice tidy format."
+ "Great that worked! We have data in a nice tidy format.\n"
]
},
{
@@ -334,7 +354,7 @@
"- There is a regularly updated list of APIs over at this [public APIs repo on github](https://github.com/public-apis/public-apis). It doesn't have an economics section (yet), but it has a LOT of other APIs.\n",
"- Berkeley Library maintains a [list of economics APIs](https://guides.lib.berkeley.edu/c.php?g=4395&p=7995952) that is well worth looking through.\n",
"- [NASDAQ Data Link](https://docs.data.nasdaq.com/), which has a great deal of [financial data](https://docs.data.nasdaq.com/docs/data-organization).\n",
- "- [DBnomics](https://db.nomics.world/): publicly-available economic data provided by national and international statistical institutions, but also by researchers and private companies."
+ "- [DBnomics](https://db.nomics.world/): publicly-available economic data provided by national and international statistical institutions, but also by researchers and private companies.\n"
]
},
{
@@ -347,7 +367,7 @@
"\n",
"Webscraping is a way of grabbing information from the internet that was intended to be displayed in a browser. But it should only be used as a last resort, and only then when permitted by the terms and conditions of a website.\n",
"\n",
- "If you're getting data from the internet, it's much better to use an API whenever you can: grabbing information in a structure way is *exactly* why APIs exist. APIs should also be more stable than websites, which may change frequently. Typically, if an organisation is happy for you to grab their data, they will have made an API expressly for that purpose. It's pretty rare that there's a major website which *does* permit webscraping but which doesn't have an API; for these websites, if they don't have an API, chances scraping is against their terms and conditions. Those terms and conditions may be enforceable by law (different rules in different countries here, and you really need legal advice if it's not unambiguous as to whether you can scrape or not.)\n",
+ "If you're getting data from the internet, it's much better to use an API whenever you can: grabbing information in a structure way is _exactly_ why APIs exist. APIs should also be more stable than websites, which may change frequently. Typically, if an organisation is happy for you to grab their data, they will have made an API expressly for that purpose. It's pretty rare that there's a major website which _does_ permit webscraping but which doesn't have an API; for these websites, if they don't have an API, chances scraping is against their terms and conditions. Those terms and conditions may be enforceable by law (different rules in different countries here, and you really need legal advice if it's not unambiguous as to whether you can scrape or not.)\n",
"\n",
"There are other reasons why webscraping is not so good; for example, if you need a back-run then it might be offered through an API but not shown on the webpage. (Or it might not be available at all, in which case it's best to get in touch with the organisation or check out WaybackMachine in case they took snapshots).\n",
"\n",
@@ -355,13 +375,13 @@
"\n",
"If you do find yourself in a scraping situation, be really sure to check that's legally allowed and also that you are not violating the website's `robots.txt` rules: this is a special file on almost every website that sets out what's fair play to crawl (conditional on legality) and what robots should not go poking around in.\n",
"\n",
- "In Python, you are spoiled for choice when it comes to webscraping. There are five very strong libraries that cover a real range of user styles and needs: **requests**, **lxml**, **beautifulsoup**, **selenium**, and *scrapy**.\n",
+ "In Python, you are spoiled for choice when it comes to webscraping. There are five very strong libraries that cover a real range of user styles and needs: **requests**, **lxml**, **beautifulsoup**, **selenium**, and \\*scrapy\\*\\*.\n",
"\n",
"For quick and simple webscraping, my usual combo would **requests**, which does little more than go and grab the HTML of a webpage, and **beautifulsoup**, which then helps you to navigate the structure of the page and pull out what you're actually interested in. For dynamic webpages that use javascript rather than just HTML, you'll need **selenium**. To scale up and hit thousands of webpages in an efficient way, you might try **scrapy**, which can work with the other tools and handle multiple sessions, and all other kinds of bells and whistles... it's actually a \"web scraping framework\".\n",
"\n",
"It's always helpful to see coding in practice, so that's what we'll do now, but note that we'll be skipping over a lot of important detail such as user agents, being 'polite' with your scraping requests, being efficient with caching and crawling.\n",
"\n",
- "In lieu of a better example, let's scrape the research page of [http://aeturrell.com/](http://aeturrell.com/)"
+ "In lieu of a better example, let's scrape the research page of [http://aeturrell.com/](http://aeturrell.com/)\n"
]
},
{
@@ -384,7 +404,7 @@
"source": [
"Okay, what just happened? We asked requests to grab the HTML of the webpage and then printed the first 300 characters of the text that it found.\n",
"\n",
- "Let's now parse this into something humans can read (or can read more easily) using beautifulsoup:"
+ "Let's now parse this into something humans can read (or can read more easily) using beautifulsoup:\n"
]
},
{
@@ -404,7 +424,7 @@
"id": "5748e928",
"metadata": {},
"source": [
- "Now we see more structure of the page and even some *HTML tags* such as 'title' and 'link'. Now we come to the data extraction part: say we want to pull out every paragraph of text, we can use beautifulsoup to skim down the HTML structure and pull out only those parts with the paragraph tag ('p').\n"
+ "Now we see more structure of the page and even some _HTML tags_ such as 'title' and 'link'. Now we come to the data extraction part: say we want to pull out every paragraph of text, we can use beautifulsoup to skim down the HTML structure and pull out only those parts with the paragraph tag ('p').\n"
]
},
{
@@ -426,7 +446,7 @@
"id": "2936677e",
"metadata": {},
"source": [
- "Although this paragraph isn't too bad, you can make this more readable by stripping out HTML tags altogether with the `.text` method:"
+ "Although this paragraph isn't too bad, you can make this more readable by stripping out HTML tags altogether with the `.text` method:\n"
]
},
{
@@ -445,7 +465,7 @@
"id": "9d9d890e",
"metadata": {},
"source": [
- "Now let's say we didn't care about most of the page, we *only* wanted to get hold of the names of projects. For this we need to identify the tag type of the element we're interested in, in this case 'div', and it's class type, in this case \"project-name\". We do it like this (and show nice text in the process):\n"
+ "Now let's say we didn't care about most of the page, we _only_ wanted to get hold of the names of projects. For this we need to identify the tag type of the element we're interested in, in this case 'div', and it's class type, in this case \"project-name\". We do it like this (and show nice text in the process):\n"
]
},
{
@@ -478,7 +498,7 @@
"info_on_pages = [scraper(root_url + str(i)) for i in range(start, stop)]\n",
"```\n",
"\n",
- "That's all we'll cover here but remember we've barely *scraped* the surface of this big, complex topic. If you want to read about an application, it's hard not to recommend the paper on webscraping that has undoubtedly change the world the most, and very likely has affected your own life in numerous ways: [\"The PageRank Citation Ranking: Bringing Order to the Web\"](http://ilpubs.stanford.edu:8090/422/) by Page, Brin, Motwani and Winograd. For a more in-depth example of webscraping, check out realpython's [tutorial](https://realpython.com/python-web-scraping-practical-introduction/)."
+ "That's all we'll cover here but remember we've barely _scraped_ the surface of this big, complex topic. If you want to read about an application, it's hard not to recommend the paper on webscraping that has undoubtedly change the world the most, and very likely has affected your own life in numerous ways: [\"The PageRank Citation Ranking: Bringing Order to the Web\"](http://ilpubs.stanford.edu:8090/422/) by Page, Brin, Motwani and Winograd. For a more in-depth example of webscraping, check out realpython's [tutorial](https://realpython.com/python-web-scraping-practical-introduction/).\n"
]
},
{
@@ -489,11 +509,11 @@
"source": [
"### Webscraping Tables\n",
"\n",
- "Often there are times when you don't actually want to scrape an entire webpage and all you want is the data from a *table* within the page. Fortunately, there is an easy way to scrape individual tables using the **pandas** package.\n",
+ "There are times when you don't need to scrape an entire webpage; you simply want the structured data from a specific table. While Polars is a high-performance data engine, it focuses on strict data formats (like Parquet or CSV) and does not natively include an HTML parser. However, we can easily bridge this gap by using Pandas to fetch the table and then converting it into a Polars DataFrame.\n",
"\n",
- "We will read data from a table on 'https://webscraper.io/test-sites/tables' using **pandas**. The function we'll use is `read_html()`, which returns a list of data frames of all the tables it finds when you pass it a URL. If you want to filter the list of tables, use the `match=` keyword argument with text that only appears in the table(s) you're interested in.\n",
+ "We will read data from 'https://webscraper.io/test-sites/tables' using `pd.read_html()`. This function scans the webpage and returns a list of all tables it finds as DataFrames. To target a specific table, we use the match= keyword argument with text that uniquely appears in the table we want—in this case, \"First Name\".\n",
"\n",
- "The example below shows how this works; looking at the website, we can see that the table we're interested in, has a 'First Name' column. Therefore we run:"
+ "Once captured, we convert the result to Polars using pl.from_pandas() to take advantage of Polars' superior query performance and expression API.\n"
]
},
{
@@ -503,10 +523,13 @@
"metadata": {},
"outputs": [],
"source": [
- "df_list = pd.read_html(\"https://webscraper.io/test-sites/tables\", match=\"First Name\")\n",
+ "import polars as pl\n",
+ "\n",
+ "pd_list = pd.read_html(\"https://webscraper.io/test-sites/tables\", match=\"First Name\")\n",
"# Retrieve first entry from list of data frames\n",
- "df = df_list[0]\n",
- "df.head()"
+ "df = pl.from_pandas(pd_list[0])\n",
+ "\n",
+ "print(df.head())"
]
},
{
@@ -515,9 +538,9 @@
"id": "31e49317",
"metadata": {},
"source": [
- "This gives us the table neatly loaded into a **pandas** data frame ready for further use.\n",
+ "This gives us the table neatly loaded into a **polars** data frame ready for further use.\n",
"\n",
- "If you get a '403' error, it means that the website has blocked **pandas** because it can see that you are engaged in web scraping. This is because some people web scrape irresponsibly, or because websites have provided other, preferred ways for you to obtain the data, eg via a download of the whole thing (think Wikipedia) or through an API. (If you really need to, [you can often get around the 403 error](https://stackoverflow.com/questions/43590153/http-error-403-forbidden-when-reading-html) though.)"
+ "If you get a '403' error, it means that the website has blocked **pandas** because it can see that you are engaged in web scraping. This is because some people web scrape irresponsibly, or because websites have provided other, preferred ways for you to obtain the data, eg via a download of the whole thing (think Wikipedia) or through an API. (If you really need to, [you can often get around the 403 error](https://stackoverflow.com/questions/43590153/http-error-403-forbidden-when-reading-html) though.)\n"
]
}
],
diff --git a/workflow-help.quarto_ipynb_1 b/workflow-help.quarto_ipynb_1
new file mode 100644
index 0000000..e7bebf8
--- /dev/null
+++ b/workflow-help.quarto_ipynb_1
@@ -0,0 +1,115 @@
+{
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Postscript: Getting Further Help {#sec-workflow-help}\n",
+ "\n",
+ "This book is not an island; there is no single resource that will allow you to master Python for Data Science. As you begin to apply the techniques described in this book to your own data, you will soon find questions that we do not answer. This section describes a few tips on how to get help, and to help you keep learning.\n",
+ "\n",
+ "## Resources\n",
+ "\n",
+ "Some other resources for learning are:\n",
+ "\n",
+ "- [The Python Data Science Handbook](https://jakevdp.github.io/PythonDataScienceHandbook/)\n",
+ "- [Real Python](https://realpython.com/), which has excellent short tutorials that cover Python more broadly (not just data science)\n",
+ "- [freeCodeCamp's Python courses](https://www.freecodecamp.org/news/search?query=data%20science%20python), though take care to select one that's at the right level for you\n",
+ "- [Coding for Economists](https://aeturrell.github.io/coding-for-economists), which has similar content to this book but is more in depth and aimed at analysts (particularly in economics)\n",
+ "\n",
+ "## Google is your friend\n",
+ "\n",
+ "If you get stuck, start with Google. Typically adding \"Python\" or \"Python Data Science\" (as the Python ecosystem goes *well* beyond data science) to a query is enough to restrict it to relevant results. Google is particularly useful for error messages. If you get an error message and you have no idea what it means, try googling it! Chances are that someone else has been confused by it in the past, and there will be help somewhere on the web.\n",
+ "\n",
+ "If Google doesn't help, try [Stack Overflow](http://stackoverflow.com). Start by spending a little time searching for an existing answer, including `[Python]` to restrict your search to questions and answers that use Python.\n",
+ "\n",
+ "## In the loop\n",
+ "\n",
+ "It's also helpful to keep an eye on the latest developments in data science. There are tons of data science newsletters out there, and we recommend keeping up with the Python data science community by following the (#pydata), (#datascience), and (#python) hashtags on Twitter.\n",
+ "\n",
+ "## Making a reprex (reproducible example)\n",
+ "\n",
+ "If your googling doesn't find anything useful, it's a really good idea prepare a minimal reproducible example or **reprex**.\n",
+ "\n",
+ "A good reprex makes it easier for other people to help you, and often you'll figure out the problem yourself in the course of making it. There are two parts to creating a reprex:\n",
+ "\n",
+ "- First, you need to make your code reproducible. This means that you need to capture everything, i.e., include any packages you used and create all necessary objects. The easiest way to make sure you've done this is to use the [**watermark**](https://github.com/rasbt/watermark) package alongside whatever else you are doing:"
+ ],
+ "id": "22b3f9e0"
+ },
+ {
+ "cell_type": "code",
+ "metadata": {},
+ "source": [
+ "import pandas as pd\n",
+ "import numpy as np\n",
+ "import watermark.watermark as watermark\n",
+ "\n",
+ "print(watermark())\n",
+ "print(watermark(iversions=True, globals_=globals()))"
+ ],
+ "id": "a119501b",
+ "execution_count": null,
+ "outputs": []
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "- Second, you need to make it minimal. Strip away everything that is not directly related to your problem. This usually involves creating a much smaller and simpler Python object than the one you're facing in real life or even using built-in data.\n",
+ "\n",
+ "That sounds like a lot of work! And it can be, but it has a great payoff:\n",
+ "\n",
+ "- 80% of the time creating an excellent reprex reveals the source of your problem. It's amazing how often the process of writing up a self-contained and minimal example allows you to answer your own question.\n",
+ "\n",
+ "- The other 20% of time you will have captured the essence of your problem in a way that is easy for others to play with. This substantially improves your chances of getting help.\n",
+ "\n",
+ "There are several things you need to include to make your example reproducible: Python environment, required packages, data, and code.\n",
+ "\n",
+ "- **Python environment**--really just the Python version. This is covered by the first call to the **watermark** package.\n",
+ "\n",
+ "- **Packages** and their versions. These should be loaded at the top of the script, so it's easy to see which ones the example needs. By using **watermark** with the above configuration, you will also print the package versions. This is a good time to check that you're using the latest version of each package; it's possible you've discovered a bug that's been fixed since you installed or last updated the package.\n",
+ "\n",
+ "- **Data**: as others won't be able to easily download the data you're working with, it's often best to create a small amount of data from code that still have the same problem as you're finding with your actual data. Between **numpy** and **pandas**, it's quite easy to generate data from code; here's an example:"
+ ],
+ "id": "c4ac60b4"
+ },
+ {
+ "cell_type": "code",
+ "metadata": {},
+ "source": [
+ "df = pd.DataFrame(\n",
+ " data=np.reshape(range(36), (6, 6)),\n",
+ " index=[\"a\", \"b\", \"c\", \"d\", \"e\", \"f\"],\n",
+ " columns=[\"col\" + str(i) for i in range(6)],\n",
+ " dtype=float,\n",
+ ")\n",
+ "df[\"random_normal\"] = np.random.normal(size=6)\n",
+ "df"
+ ],
+ "id": "d1e4562c",
+ "execution_count": null,
+ "outputs": []
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "- **Code**: copy and paste the minimal reproducible example code (including the packages, as noted above). Make sure you've used spaces and your variable names are concise, yet informative. Use comments to indicate where your problem lies. Do your best to remove everything that is not related to the problem. Finally, the shorter your code is, the easier it is to understand, and the easier it is to fix.\n",
+ "\n",
+ "Finish by checking that you have actually made a reproducible example by starting a fresh Python session and copying and pasting your reprex in."
+ ],
+ "id": "4b75e409"
+ }
+ ],
+ "metadata": {
+ "kernelspec": {
+ "name": "python3",
+ "language": "python",
+ "display_name": "Python 3 (ipykernel)",
+ "path": "/Users/omagic/Documents/GitHub/python4DSpolars/.venv/share/jupyter/kernels/python3"
+ }
+ },
+ "nbformat": 4,
+ "nbformat_minor": 5
+}
\ No newline at end of file
diff --git a/workflow-packages-and-environments.quarto_ipynb_1 b/workflow-packages-and-environments.quarto_ipynb_1
new file mode 100644
index 0000000..a5600ce
--- /dev/null
+++ b/workflow-packages-and-environments.quarto_ipynb_1
@@ -0,0 +1,149 @@
+{
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Workflow: Packages and Environments {#sec-workflow-packages-and-environments}\n",
+ "\n",
+ "In this chapter, you're going to learn about packages and how to install them plus virtual coding environments that keep your packages isolated and your projects reproducible.\n",
+ "\n",
+ "## Packages\n",
+ "\n",
+ "### Introduction\n",
+ "\n",
+ "Packages (also called libraries) are key to extending the functionality of Python. It won't be long before you'll need to install some. There are packages for geoscience, for building websites, for analysing genetic data, for economics—pretty much for anything you can think of. Packages are typically not written by the core maintainers of the Python language but by enthusiasts, firms, researchers, academics, all sorts! Because anyone can write packages, they vary widely in their quality and usefulness. There are some that you'll be seeing them again and again.\n",
+ "\n",
+ "
\n",
+ "\n",
+ "The three Python packages **numpy**, **pandas**, and **maplotlib**, which respectively provide numerical, data analysis, and plotting functionality, are ubiquitous. So many scripts begin by importing all three of them, as in the tweet above!\n",
+ "\n",
+ "There are typically two steps to using a new Python package:\n",
+ "\n",
+ "1. *install* the package on the command line (aka the terminal), eg using `uv add pandas`\n",
+ "\n",
+ "2. *import* the package into your Python session, eg using `import pandas as pd`\n",
+ "\n",
+ "When you issue an install command for a specific package, it is automatically downloaded from the internet and installed in the appropriate place on your computer. To install extra Python packages, you issue install commands to a text-based window called the \"terminal\".\n",
+ "\n",
+ "### The Command Line in Brief\n",
+ "\n",
+ "The *terminal* or *command line* or sometimes the *command prompt* was labelled 4 in the screenshot of Visual Studio Code from the chapter on @sec-introduction. The terminal is a text-based way to issue all kinds of commands to your computer (not just Python commands) and knowing a little bit about it is really useful for coding (and more) because managing packages, environments (which we haven't yet discussed), and version control (ditto) can all be done via the terminal. We'll come to these in due course in the chapter on @sec-command-line, but for now, a little background on what the terminal is and what it does.\n",
+ "\n",
+ "::: {.callout-note}\n",
+ "To open up the command line within Visual Studio Code, use the ⌃ + \\` keyboard shortcut (Mac) or ctrl + \\` (Windows/Linux), or click \"View > Terminal\".\n",
+ "\n",
+ "If you want to open up the command line independently of Visual Studio Code, search for \"Terminal\" on Mac and Linux, and \"Powershell\" on Windows.\n",
+ ":::\n",
+ "\n",
+ "Firstly, everything you can do by clicking on icons to launch programmes on your computer, you can also do via the terminal, also known as the command line. For many programmes, a lot of their functionality can be accessed using the command line, and other programmes *only* have a command line interface (CLI), including some that are used for data science.\n",
+ "\n",
+ "::: {.callout-tip}\n",
+ "The command line interacts with your operating system and is used to create, activate, or change Python installations.\n",
+ ":::\n",
+ "\n",
+ "Use Visual Studio Code to open a terminal window by clicking Terminal -> New Terminal on the list of commands at the very top of the window. If you have installed uv on your computer, your terminal should look something like this as your 'command prompt':\n",
+ "\n",
+ "```bash\n",
+ "your-username@your-computer current-directory %\n",
+ "```\n",
+ "\n",
+ "on Mac, and the same but with '%' replaced by '$' on linux, and (using Powershell)\n",
+ "\n",
+ "```powershell\n",
+ "PS C:\\Windows\\System32>\n",
+ "```\n",
+ "\n",
+ "on Windows.\n",
+ "\n",
+ "You can check that uv has successfully installed Python in your current project's folder by running\n",
+ "\n",
+ "```bash\n",
+ "uv run python --version\n",
+ "```\n",
+ "\n",
+ "For now, to at least try out the command line, let's use something that works across all three of the major operating systems. Type `uv run python` on the command prompt that came up in your new terminal window. You should see information about your installation of Python appear, including the version, followed by a Python prompt that looks like `>>>`. This is a kind of interactive Python session, in the terminal. It's much less rich than the one available in Visual Studio Code (it can't run scripts line-by-line, for example) but you can try `print('Hello World!')` and it will run, printing your message. To exit the terminal-based Python session, type `exit()` to go back to the regular command line.\n",
+ "\n",
+ "### Installing Packages\n",
+ "\n",
+ "To install extra Python packages, the default and easiest way is to use `uv add **packagename**`. There are over 330,000 Python packages on PyPI (the Python Package Index)! You can see what packages you have installed already by running `uv pip list` into the command line.\n",
+ "\n",
+ "`uv add ...` will install packages into the special Python environment in your current folder (it sits in a subdirectory called \".venv\" which will be hidden by default on most systems.) It's really helpful and good practice to have one Python environment per project, and **uv** does this automatically for you.\n",
+ "\n",
+ "::: {.callout-tip title=\"Exercise\"}\n",
+ "Try installing the **matplotlib**, **pandas**, **statsmodels**, and **skimpy** packages using `uv add`.\n",
+ ":::\n",
+ "\n",
+ "### Using Packages\n",
+ "\n",
+ "Once you have installed a package, you need to be able to use it! This is usually done via an import statement at the top of your script or Jupyter Notebook. For example, to bring in **pandas**, it's\n",
+ "\n",
+ "```python\n",
+ "import pandas as pd\n",
+ "```\n",
+ "\n",
+ "Why does Python do this? The idea of not just loading every package is to provide clarity over what function is being called from what package. It's also not necessary to load every package for every piece of analysis, and you often actually want to know what the *minimum* set of packages is to reproduce an analysis. Making the package imports explicit helps with all of that.\n",
+ "\n",
+ "You may also wonder why one doesn't just use `import pandas as pandas`. There's actually nothing stopping you doing this except i) it's convenient to have a shorter name and ii) there does tend to be a convention around imports, ie `pd` for **pandas** and `np` for **numpy**, and your code will be clearer to yourself and others if you follow the conventions.\n",
+ "\n",
+ "## Virtual Code Environments\n",
+ "\n",
+ "Virtual code environments allow you to isolate all of the packages that you're using to do analysis for one project from the set of packages you might need for a different project. They're an important part of creating a reproducible analytical pipeline but a key benefit is that others can reproduce the environment you used and it's best practice to have an isolated environment per project.\n",
+ "\n",
+ "To be more concrete, let's say you're using Python 3.9, **statsmodels**, and **pandas** for one project, project A. And, for project B, you need to use Python 3.10 with **numpy** and **scikit-learn**. Even with the same version of Python, best practice would be to have two separate virtual Python environments: environment A, with everything needed for project A, and environment B, with everything needed for project B. For the case where you're using different versions of Python, this isn't just best practice, it's essential.\n",
+ "\n",
+ "Many programming languages now come with an option to install packages and a version of the language in isolated environments. In Python, there are multiple tools for managing different environments. And, of those, the easiest to work with is probably [**uv**](https://docs.astral.sh/uv/).\n",
+ "\n",
+ "You can see all of the packages in the environment created in your current folder by running `uv pip list` on the command line. Here's an example of looking at the installed packages within this very book, filtering them just to the ones beginning with \"s\".\n",
+ "\n",
+ "```{bash}\n",
+ "uv run pip list | grep ^s\n",
+ "```\n",
+ "\n",
+ "### The pyproject.toml file in Python Environments\n",
+ "\n",
+ "You may have noticed that a file called `pyproject.toml` has been created."
+ ],
+ "id": "8b889898"
+ },
+ {
+ "cell_type": "code",
+ "metadata": {},
+ "source": [
+ "import toml\n",
+ "from rich import print_json\n",
+ "\n",
+ "print_json(data=toml.load(\"pyproject.toml\"))"
+ ],
+ "id": "688f09f1",
+ "execution_count": null,
+ "outputs": []
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "This lists all of the dependencies, and the version, of a **uv** Python project. There are lots of benefits to tracking what versions of packages you're using like this. One of the most important is that you can *share* projects with other people, and they can install them from these files too.\n",
+ "\n",
+ "As you install or remove packages, the `pyproject.toml` file changes in lockstep.\n",
+ "\n",
+ "Noe that Visual Studio Code shows which Python environment you are using when you open a Python script or Jupyter Notebook.\n",
+ "\n",
+ "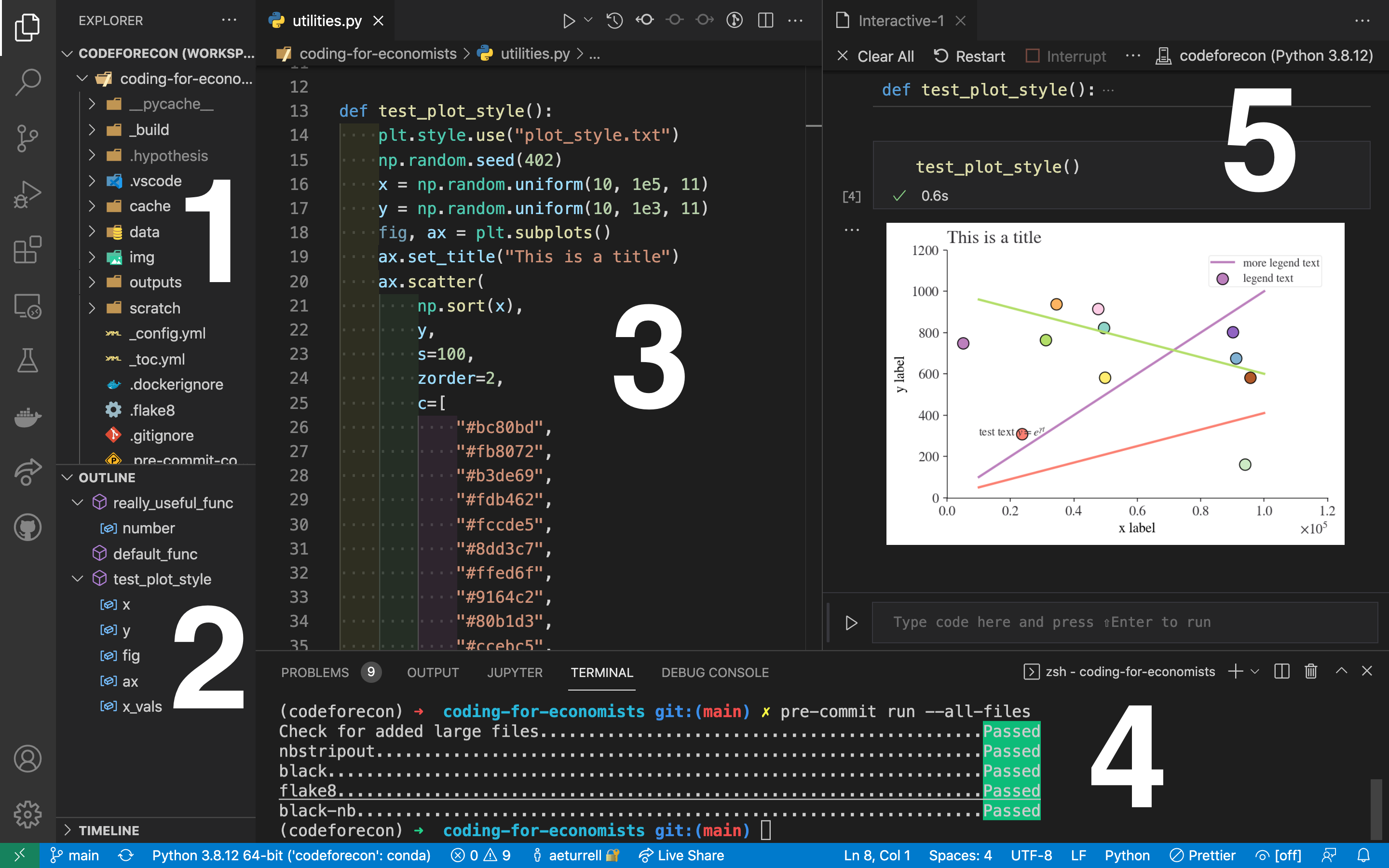\n",
+ "\n",
+ "In the screenshot above, you can see the project-environment in two places: on the blue bar at the bottom of the screen, and (in 5), at the top right hand side of the interactive window. A similar top right indicator is present when you have a Jupyter Notebook open too."
+ ],
+ "id": "148595b3"
+ }
+ ],
+ "metadata": {
+ "kernelspec": {
+ "name": "python3",
+ "language": "python",
+ "display_name": "Python 3 (ipykernel)",
+ "path": "/Users/omagic/Documents/GitHub/python4DSpolars/.venv/share/jupyter/kernels/python3"
+ }
+ },
+ "nbformat": 4,
+ "nbformat_minor": 5
+}
\ No newline at end of file